Unleashing the Power of Data, Collaboration, and Innovation in Drug Development
Pooja Arora, Principal Consultant, Thoughtworks
Drug development is one of the most complex and costly undertakings in modern healthcare. Despite the vast amount of data generated at every stage—discovery, clinical trials, regulatory review, and post-market surveillance—fragmentation and siloed practices prevent organisations from realising the full value of this information. This article explores how siloed data hampers progress, highlights scenarios where integration is critical, and outlines strategies to weave data, collaboration, and innovation into a connected knowledge fabric. The goal is clear: improving patient outcomes by accelerating safer, more effective therapies to market.
Introduction
“We don't have to look for model organisms, we are model organisms.”
— Sydney Brenner, Nobel Laureate
Drug discovery and development is one of the most data-intensive enterprises in modern science. Vast amounts of information are generated at every stage—genomic datasets, high-throughput screening results, clinical trial outcomes, regulatory submissions, real-world evidence, and patient-reported data. Yet, despite this abundance, knowledge gaps persist. Information often remains siloed, poorly connected, or described in inconsistent ways, preventing organisations from realising its full value.
Drug development also remains fraught with risk. Clinical trials, the most visible and costly stage of the process, face a staggering failure rate, with nearly 90% of drug candidates failing to progress. Many of these failures can be traced to the translational gap: promising results in preclinical animal models that do not replicate in human populations. The industry is therefore shifting from relying solely on model organisms to incorporating data from real human populations earlier in the process, creating a stronger connection between clinical (bedside) data and early discovery (bench) science.
This shift underscores the need for reverse translation—not simply moving discoveries from bench to bedside but also moving knowledge from bedside back to bench. Clinical and real-world insights must guide upstream research, enabling scientists to refine hypotheses, validate targets, and design smarter trials that reflect real patient diversity and complexity.
These challenges are not confined to clinical trials. Across the value chain, similar disconnects occur:
- In early discovery, real-world evidence could help scientists prioritise targets that are most clinically relevant, focusing scarce resources where they are most likely to yield benefit.
- In clinical development, learnings from prior studies could inform more efficient trial design, patient stratification, and endpoint selection.
- In commercialisation and delivery, trial data and post-market insights could guide patient engagement strategies, adherence programs, and precision medicine approaches tailored to subgroups.
At the root of these missed opportunities lies a common barrier: fragmented and disconnected data. Diverse nomenclature, inconsistent ontologies, and siloed systems make it difficult to integrate information across stages of development. Overcoming this requires a unifying approach—one that treats data not as isolated repositories but as part of a connected fabric.
Semantics provides the connective tissue drug development has been missing. Through ontologies, knowledge graphs, and AI-driven agents, disparate data sources can be unified into actionable insight—transforming complexity into clarity and accelerating the journey from science to patient care.
From Bottlenecks to Breakthroughs: Why We Need a Semantic Agent Framework
The future of drug development lies in transforming fragmented data into a living knowledge fabric that spans the entire value chain. This vision moves beyond incremental efficiencies to create a holistic, patient-centered ecosystem where insights continuously flow across discovery, development, and delivery.
Today’s bottlenecks are the result of siloed practices:
- Discovery scientists may lack access to clinical outcomes that could validate targets.
- Trial teams often struggle to extract safety insights from lengthy, unstructured reports.
- Pharmacovigilance functions identify safety signals only months after patient exposure.
- Commercial teams see only fragments of the patient journey, making it difficult to support adherence or personalize therapies.
Each of these issues is symptomatic of the same root cause: the industry’s inability to represent and connect knowledge in a consistent, reusable way.
A semantic agent framework addresses this gap. It unites foundational data with a semantic layer of meaning and context and then applies AI agents to orchestrate knowledge across the value chain.
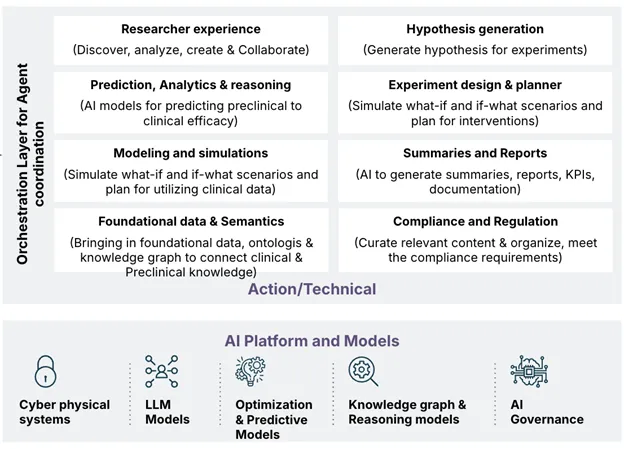
Fig 1: Representing the AI Platform & models and the different orchestration agents from foundational data to researcher experience.
The vision is a system where every piece of data, from the earliest discovery to the latest post-market report, is connected and meaningful. Rather than static repositories, data becomes part of a dynamic, living ecosystem.
At its core, this vision rests on three interconnected layers:
1. Foundational Data
- Data must be standardised, interoperable, and FAIR (Findable, Accessible, Interoperable, Reusable).
- Instead of siloed repositories, organisations need shared, reusable data assets that serve as the foundation for collaboration and innovation.
2. Semantic Layer
- Ontologies provide a common vocabulary to harmonise terms across systems.
- Knowledge graphs weave entities—genes, diseases, patients, and outcomes—into rich networks of meaning.
- This semantic structure enables reverse translation, where patient and trial insights feed upstream into discovery.
3. AI Agents and Orchestration
- AI agents act as intelligent copilots, navigating knowledge graphs, extracting insights, and recommending experiments.
- In self-driving labs, agents continuously adapt experiments based on real-world evidence and clinical outcomes, creating a perpetual learning loop.
The outcome is a feedback-rich ecosystem where real-world evidence informs early discovery, clinical trial learnings shape later-stage development and delivery, and post-market signals guide the design of next-generation therapies. In this model, every stage of the value chain reinforces the others, accelerating development while improving safety and patient outcomes.
Semantic Agent Capabilities in Action
Overcoming fragmentation requires more than better storage or faster computing. What’s needed is a framework that brings meaning, context, and intelligence into the heart of drug development. The semantic agent framework delivers this through three core capabilities:
1. Ontologies: Creating a Common Vocabulary
Biomedical concepts are often described inconsistently—“myocardial infarction” may appear as “heart attack,” “MI,” or an ICD code. Ontologies resolve this by standardising meaning across datasets. This consistency ensures that discovery teams, trial designers, regulators, and commercial functions can interpret and integrate information without losing context.
2. Knowledge Graphs: Connecting Relationships
While ontologies create clarity, knowledge graphs provide context. They link standardised concepts into networks of relationships—connecting genes, diseases, pathways, biomarkers, trial outcomes, and real-world data. Unlike static databases, knowledge graphs are dynamic, allowing researchers to ask new questions such as:
- Which patient subgroups are most likely to respond to this drug class?
- How do prior trial outcomes connect to ongoing studies?
- What links exist between real-world adverse events and molecular pathways?
By embedding relationships, knowledge graphs create the backbone for translation and reverse translation across the value chain.
3. AI Agents: Orchestrating Discovery
Once data is standardised and connected, AI agents make it actionable. Acting as copilots, they:
- Traverse knowledge graphs to surface hidden insights.
- Extract meaning from unstructured trial documents or regulatory submissions.
- Recommend next experiments or adjustments to trial design.
- Adapt preclinical pipelines dynamically in self-driving labs, where real-world patient signals guide experiments in real time.
Together, these capabilities transform drug development into a continuous learning loop, where knowledge flows seamlessly across discovery, trials, regulation, and patient care.
Outcome: Connected Entities & Enhanced Relationships
When these capabilities converge, the result is more than technical integration—it is a continuous learning loop for discovery and development. Clinical outcomes guide upstream research, safety signals are detected proactively rather than reactively, and trials are designed around real-world patient needs. Collaboration strengthens as scientists, clinicians, regulators, and medical affairs share a common knowledge fabric.
Instead of functioning as a linear relay race, drug development becomes a continuous learning system—one that adapts, evolves, and improves with every new data point.
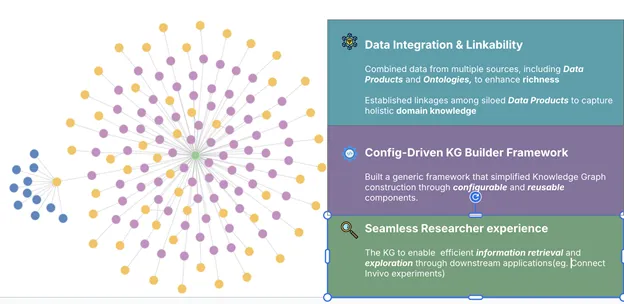
Conclusion: Connected Researcher Experience & Patient Journey
“Solving a problem simply means representing it so as to make the solution transparent.” Herbert Simon, Nobel Laureate and AI pioneer
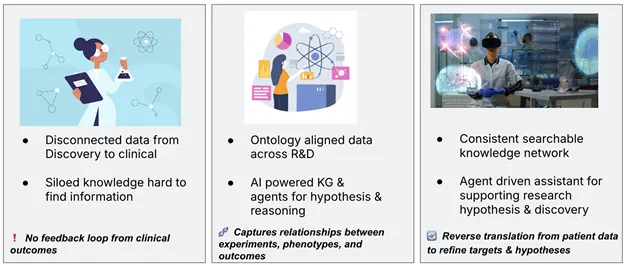
Drug development’s grand challenge is not the absence of data, but the inability to connect and represent it in ways that reveal solutions transparently. A semantic agent framework does exactly this: it represents biomedical knowledge in context, connects the dots across the value chain, and enables intelligent agents to act on insights.
For researchers, this creates an environment where data is accessible, interoperable, and collaborative. For patients, it means faster access to safer, more effective therapies tailored to their needs.
Ultimately, connecting knowledge is not just a technical fix—it is a moral imperative. By weaving together data, collaboration, and innovation, we create a future where drug development is guided not by fragmented silos but by a shared mission: improving lives.
References
- Ashburner, M., et al. (2000). "Gene ontology: tool for the unification of biology." Nature Genetics.
- Hastings, J., et al. (2016). "ChEBI in 2016: Improved services and an expanding collection of metabolites." Nucleic Acids Research.
- Schriml, L. M., et al. (2019). "Human Disease Ontology 2018 update: classification, content, and workflow expansion." Nucleic Acids Research.
- Mungall, C. J., et al. (2017). "Uberon, an integrative multi-species anatomy ontology." Genome Biology.
- Smith, B., et al. (2020). "Basic Formal Ontology 2.0: A reference ontology for biomedical informatics." Journal of Biomedical Semantics.
- Wang, Q., et al. (2018). "Knowledge Graph Embedding: A Survey of Approaches and Applications." IEEE Transactions on Knowledge and Data Engineering.
- Groth, P., & Cox, S. J. (2017). "Semantic integration for large-scale scientific data." Data Intelligence.
- Nickel, M., et al. (2015). "A review of relational machine learning for knowledge graphs." Proceedings of the IEEE.
- Brown, T. B., et al. (2020). "Language Models are Few-Shot Learners." Advances in Neural Information Processing Systems.
- Stein, L. D., et al. (2021). "The ecosystem of linked biomedical data." Nature Reviews Genetics.
- Noy, N. F., et al. (2019). "Ontology development 101: A guide to creating your first ontology." Stanford Knowledge Systems Lab.
- Loshin, D. (2012). "Big data analytics: from strategic planning to enterprise integration." Morgan Kaufmann.
- Hay, M., Thomas, D., & Leahy, L. (2014). Clinical Development Success Rates 2006-2015. Biopharmaceutical Research & Development. Nature Reviews Drug Discovery
- Shakhnovich V. (2018). It's Time to Reverse our Thinking: The Reverse Translation Research Paradigm. Clinical and translational science, 11(2), 98–99. https://doi.org/10.1111/cts.12538










