Predicting Unbound Drug Bioavailability in The Brain
Application of machine learning
Alan Talevi, Full Professor, University of La Plata, and a Principal Researcher, Argentinean Council of Science (CONICET); CEO, Boolzi.
Unbound brain drug bioavailability became a major driver of central nervous system drug discovery. Here, we discuss the potential and current limitations of machine learning applications to predict unbound brain bioavailability, and whether the major issue of scarcity of high-quality data is likely to be solved in the coming years.
In the last fifteen years, unbound brain bioavailability has become a major driver of central nervous system (CNS) drug discovery. An article from 2022 in Pharmaceutical Research by Loryan et al. describes the partition coefficient of unbound drug concentrations between the brain and plasma (Kp,uu,br) as a “game-changing parameter” in the field of CNS therapeutics. A survey published in that same article, administered to 14 representatives of major pharmaceutical companies (13 of whom belonged to their companies' DMPK department) confirmed that almost all companies have integrated the estimation of unbound brain bioavailability to establish in vitro-in vivo correlations, PK/PD relationship for CNS effects, and/or for prediction of therapeutic dose for brain medications under development.

However, the most commonly used in vivo and in vitro methods to measure free brain bioavailability (brain/blood microdialysis, brain/blood sampling followed by binding correction, the brain slices assay, and the brain homogenate assay) present various limitations, from systematic errors in the determination (e.g., brain homogenate assay) to high cost and need for highly trained personnel (e.g., brain microdialysis). The greatest of these limitations is, possibly, their throughput, which is mid-to-high at best.
This possibly explains why more than half of companies resort to QSAR or physiologically based pharmacokinetic modelling (PBPK) models to determine brain tissue binding and uptake, and to predict the distribution of unbound drugs. Once such models are built, they can be applied with high throughput at no cost. Moreover, experts point out the establishment of a truly predictable QSAR model as one of the key developments for the implementation of the unbound brain bioavailability concept in the next 15 years.
Without a doubt, today we have machine learning tools capable of developing highly predictive models to predict unbound brain bioavailability. Considerable efforts have been made recently to obtain such models (including applications of a variety of algorithms that include neural networks, ensemble learning, random forests, support vector machines, gradient boosting, and also physics-based approaches). The best of them have achieved, at best, a moderate predictive ability if judged from their internal and/or external determination coefficients, in the case of regression models, or the overall accuracy, in the case of classification models (see Table 1 for a summary of some recent reports). To give an example, the accuracy of the reported classifier models hardly exceeds 85 per cent, which is good, but far from the real possibilities of artificial intelligence today. (Table 1)
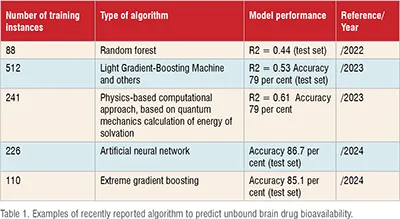
So, what is happening? Why aren't we exploiting the full potential of machine learning to predict unbound brain drug bioavailability (a question that could be extended to many other pharmaceutically relevant properties.)? Why are state-of-the-art algorithms not being fully exploited to obtain the highest possible accuracy in predictions? It has been established that one of the keys to developing global (universal) models with increased predictive capacity is the availability of larger datasets with high quality.
Unbound drug bioavailability in the brain depends on the complex interplay between multiple factors, including passive permeability, facilitated transport, and active uptake and efflux transport by a diversity of drug transporters at the blood-brain barrier. Opportunely, the latest generation of machine learning algorithms have been conceived to deal with complex problems much more challenging than the chemical-mathematical relationships between the molecular structure of small or large molecules and their physicochemical, biological, or pharmacological properties. Still, these powerful algorithms are prone to overfitting if applied to relatively small data sets, and doomed to failure if such data are not of high quality (under the well-known principle of "garbage in, garbage out"). The availability of large-scale data is thus a precondition for the application of highly flexible algorithms.
It is likely that this landscape of data scarcity will change profoundly (for the better) in the coming years. The pharmaceutical industry and other relevant players in the field of biomedicine have increasingly recognised the acceleration potential of collaborative data initiatives, which also greatly favour reproducibility in research. The synergistic potential of the open knowledge paradigm and data democratisation is widely recognised in the world of computing, where the open-source philosophy began. Such a philosophy largely explains the current revolution in the field of communication technologies, embodied today in highimpact initiatives promoted by giants such as Microsoft, Google, or OpenAI, among which could be mentioned Azure OpenAI, GitHub, GitHub Copilot or, well within the field biomedical sciences, AlphaFold. In view of the impressive results obtained, this collaborative spirit will likely spread to other areas in the coming years.
Interestingly, while security and privacy of shared clinical/medical data remains paramount within pharmaceutical companies to ensure compliance with regulatory standards, it is conceivable that restrictions and precautions could be relaxed significantly in the case of shared preclinical data, such as unbound drug bioavailability values for proprietary compounds. And that the cost-benefit balance of sharing a small selection of data acquired in-house is such that the benefits far outweigh the imaginable losses.
The above would solve the issue of data scarcity. What about the quality of that data? The quality of the data will improve as the experimental techniques used to estimate unbound brain bioavailability are perfected and best practices are implemented, minimising the systematic errors and inter-assay variation inherent to each of them. A good example is the use of anti-inflammatory agents and microfabricated ultrasmall probes in brain microdialysis experiments to better preserve the blood-brain barrier functionality upon implantation of the probe.
References:
1. Loryan I, Reichel A, Feng B, Bundgaard C, Shaffer C, Kalvass C, Bednarczyk D, Morrison D, Lesuisse D, Hoppe E, Terstappen GC, Fischer H, Di L, Colclough N, Summerfield S, Buckley ST, Maurer TS, Fridén M. Unbound brain-to-plasma partition coefficient, Kp,uu,brain-a game changing parameter for CNS drug discovery and development. Pharm Res. 2022;39(7):1321-1341. doi: 10.1007/s11095-022-03246-6.
2. Fridén M, Gupta A, Antonsson M, Bredberg U, Hammarlund-Udenaes M. In vitro methods for estimating unbound drug concentrations in the brain interstitial and intracellular fluids. Drug Metab Dispos. 2007; 35(9):1711-9. doi: 10.1124/dmd.107.015222.
3. Talevi A (editor). CNS Drug Development and Delivery Concepts and Applications. Springer Cham, 2024.
4. Ma Y, Jiang M, Javeria H, Tian D, Du Z. Accurate prediction of Kp,uu,brain based on experimental measurement of Kp,brain and computed physicochemical properties of candidate compounds in CNS drug discovery. Heliyon. 2024; 10(2):e24304. doi: 10.1016/j.heliyon.2024.e24304.
5. Morales JF, Ruiz ME, Stratford RE, Talevi A. Application of machine learning to predict unbound drug bioavailability in the brain. Front Drug Discov. 2024; 4: 1360732. doi: 10.3389/fddsv.2024.1360732.
6. Umemori Y, Handa K, Sakamoto S, Kageyama M, Iijima T. QSAR model to predict Kp,uu,brain with a small dataset, incorporating predicted values of related parameter. SAR QSAR Environ Res. 2022; 33(11):885-897. doi: 10.1080/1062936X.2022.2149619.
7. Liu S, Kosugi Y. Human brain penetration prediction using scaling approach from animal machine learning models. AAPS J. 2023; 25(5):86. doi: 10.1208/s12248-023-00850-1. PMID: 37667061,
8. Lawrenz M, Svensson M, Kato M, Dingley KH, Chief Elk J, Nie Z, Zou Y, Kaplan Z, Lagiakos HR, Igawa H, Therrien E. A computational physics-based approach to predict unbound brain-to-plasma partition coefficient, Kp,uu. J Chem Inf Model. 2023; 63(12):3786-3798. doi: 10.1021/acs.jcim.3c00150.
9. Liu H, Dong K, Zhang W, Summerfield SG, Terstappen GC. Prediction of brain:blood unbound concentration ratios in CNS drug discovery employing in silico and in vitro model systems. Drug Discov Today. 2018;23(7):1357-1372. doi: 10.1016/j.drudis.2018.03.002.
10. Ruatta SM, Prada Gori DN, Fló Díaz M, Lorenzelli F, Perelmuter K, Alberca LN, Bellera CL, Medeiros A, López GV, Ingold M, Porcal W, Dibello E, Ihnatenko I, Kunick C, Incerti M, Luzardo M, Colobbio M, Ramos JC, Manta E, Minini L, Lavaggi ML, Hernández P, Šarlauskas J, Huerta García CS, Castillo R, Hernández-Campos A, Ribaudo G, Zagotto G, Carlucci R, Medrán NS, Labadie GR, Martinez-Amezaga M, Delpiccolo CML, Mata EG, Scarone L, Posada L, Serra G, Calogeropoulou T, Prousis K, Detsi A, Cabrera M, Alvarez G, Aicardo A, Araújo V, Chavarría C, Mašič LP, Gantner ME, Llanos MA, Rodríguez S, Gavernet L, Park S, Heo J, Lee H, Paul Park KH, Bollati-Fogolín M, Pritsch O, Shum D, Talevi A, Comini MA. Garbage in, garbage out: how reliable training data improved a virtual screening approach against SARS-CoV-2 MPro. Front Pharmacol. 2023; 14:1193282. doi: 10.3389/fphar.2023.1193282.
11. Nesbitt KM, Varner EL, Jaquins-Gerstl A, Michael AC. Microdialysis in the rat striatum: effects of 24 h dexamethasone retrodialysis on evoked dopamine release and penetration injury. ACS Chem Neurosci. 2015;6(1):163-73. doi: 10.1021/cn500257x.
12. Ngernsutivorakul T, Steyer DJ, Valenta AC, Kennedy RT. In vivo chemical monitoring at high spatiotemporal resolution using microfabricated sampling probes and droplet-based microfluidics coupled to mass spectrometry. Anal Chem. 2018; 90(18):10943-10950. doi: 10.1021/acs.analchem.8b02468.











