Personalised Healthcare
Hitting the mark
Shannon A Graver, Global Studies Operations Manager, PDOC
F Hoffmann-La Roche, Switzerland
The concept of Personalised Healthcare (PHC) is being driven by the idea of improved patient outcomes and also to contain soaring healthcare costs. It is only when the right patients receive the right treatment that the true value of PHC is realised. To achieve this, biomarkers that are quantifiable need to be identified to select the right patient populations. The challenges of implementing this new research strategy are complex and would require a multifaceted approach.
Personalised Healthcare is not a novel concept, but has undergone a continual evolution. More than 1000 years ago diagnosis and treatments were based on what could be seen, smelt, tasted, palpated or intuited. About 100 years ago, diagnosis and treatment started to be based on a greater understanding of surgery, biochemistry and cellular processes. Today, the focus is getting sharper and rapidly growing insights into molecular processes and variations in our genes, is changing the path of medicine.
The aims of Personalised Healthcare (PHC) initiatives are simple: only treat those patients who are likely to respond to a given treatment. By achieving this, the efficacy and safety for every patient can be greatly improved. However, the reality is, that science and medical advancements have been sluggish to capitalise.
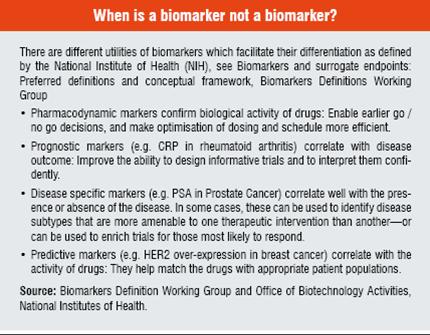
The term “Biomarker” is all encompassing; it requires our understanding of drug metabolism, action, efficacy and / or safety; facilitates prediction of the response to therapies; can expand the molecular definition of disease; and can recognise the stage of diseases. According to this broad definition, biomarkers include all diagnostic tests, imaging technologies and any other objective measure of a person’s health status and all pharmacodiagnostic tests. Genetics, genomics, proteomics, modern imaging techniques and other technologies enable this detection and quantification of many more markers than ever before which leads to an improvement in our understanding of targets, signaling pathways, metabolism and mechanisms of toxicity.
Today molecular technologies are available to better target treatments, but discovering and developing novel medicines and diagnostic markers (biomarkers) is a very complex and time-consuming undertaking. By evaluating the individual characteristics of patients and their diseases (e.g. cancer subtypes) into account, PHC could
1. improve medical outcomes and the quality of care;
2. predict which patients will most likely benefit—helping avoid treatment without benefit;
3. aid in the development of safer and more effective treatments by reducing the risk of side effects; and
4. save patients’ lives and improve their quality of life.
By increasing efficacy and safety, PHC could
1. make therapies more cost-effective;
2. create diagnostic products that can help save costs by targeting therapies to the patients who are most likely to respond;
3. create diagnostic products may also help avoid severe side effects
4. make more efficient use of healthcare budgets.
“Omics”
The Human Genome Project (HGP) took 13 years to be completed (2003). The project identified the human DNA was made up of approximately 3 billion chemical base pairs and 25,000-30,000 genes. As a result, genetic strategies were the first to adopt the concept of personalised healthcare, and the pharmacogenomics era was born. Pharmacogenomics is an example of revolutionary technologies and evolutionary practices coming together to determine an individual response to drugs that is based on the affects of genetic inheritance as opposed to the traditional means of a “one size fits all” approach. The benefits of a diagnosis based on genotypic and integrated phenotypic data could result in significant improvements in earlier treatments and extending not only the life of patient populations but also the quality. However, the limitations of pharmacogenomics soon became apparent as it failed to account for individual environmental factors. As a consequence, and as an alternative approach to pharmacogenomics, research into the metabolic profile, the so called pharmacometabonomics, was undertaken. This technique could demonstrate for example that pre-dose urinary profiles carried information about the degree of toxicity post-dosing and, in the case of paracetamol, information that is predictive of the drug’s in vivo biotransformation. Furthermore, a responder / non-responder pattern of liver damage at 24 hour post-dosing was reflected in the pre-dose metabolic profile of the urine, as described by Nicholson JK, Molecular Systems Biology 2:52. These studies show that there is a realistic and valuable approach to screen human populations with plasma metabolic profiles which result in indicative measures.
Given the reality that therapeutics appear to be effective for only 20 to 60 per cent of patients prescribed, and, nearly 200,000 people die from adverse drug reactions each year, researchers and physicians need new tools to be able to combat the variability of diseases. Within the next decade, the development of molecular diagnostic products will most likely give researchers the tools to predict a patient’s therapeutic response that is based on either the patient’s inherited genetics, or the genetic makeup of a tumour or the viral genotype.
If it were not for the great variability among individuals, medicine might as well be a science and not an art.
– Sir William Osler
A biomarker is a characteristic that is objectively measured and evaluated as an indicator of normal biological processes, pathogenic processes or pharmacological responses to therapeutic intervention, as defined by the biomarker definition working group, Clinical Pharmacology & Therapeutics (2001) 69, 89–95. Some unanswered questions where the solution could hide in the discovery of novel markers include, the prevalence of diabetes that is highest among Alaskan Natives and American Indians; and that men are more likely to die from congestive heart disease than women.
Biomarkers may or may not be dynamically modulated (e.g. DNA, which generally does not change, versus metabolites or proteins, which do change). Medicine and science are moving in this direction, as some of the following examples might tell: We measure blood glucose to tailor insulin treatment to patients’ needs. In the case of Osteoporosis, a broad range of tests is available to assess bone integrity and to monitor the effects of antiresorptive therapy. Patients suffering from HIV are being tested in order to measure viral levels in HIV patients before and during treatment with an antiretroviral drug, this allows physicians to monitor success as well as evolving resistance to the therapy.
In the large area of oncology, breast cancer is a perfect example. It is globally accepted to measure the presence / expression of a growth factor (HER2) in breast cancer with specific tests such as IHC, FISH or CISH, identifying patients who are likely to respond to Herceptin, a therapy that specifically targets this growth factor. Another important case for patients with Chronic Myelogenous Leukaemia (CML) is Gleevec where patients being tested for the presence of mutations in BCR-ABL are also used to monitor drug resistance to the treatment.
However, for predicting individual drug response, the gene-chip technology, AmpliChip CYP450 test, the world’s first commercial pharmacogenetic product, analyses variations in two genes that play a major role in the metabolism of many widely prescribed drugs. Another example of current advances in molecular diagnostics is a myocardial infarction diagnostic tool, developed by deCODE genetics and Affymetrix which is a prognostic test that gives insight into the probability of patients developing a myocardial infarction.
There are many areas where progress is being made by developing a sound biomarker strategy. It has been shown recently that biomarkers are unlocking some indicative mechanisms such as the inhibition of tyrosine kinase in tumours with EGFR somatic mutations; the influences on the response to Gefitinib, Irinotecan with UGT1A1 mutations; Warfarin with CYP2C9, and VKORC1 and DPD polymorphisms and ability to metabolise 5FU
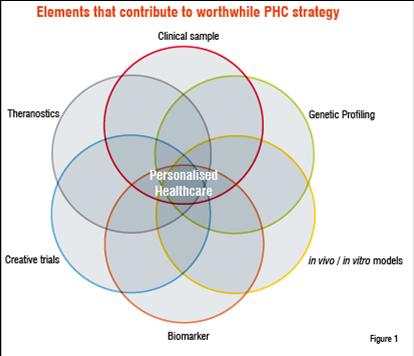
Novel molecular and imaging technologies, such as genetics, genomics, proteomics and PET / DCE-MRI imaging are increasingly used in early drug development to try and identify biomarkers which may enable advancements in PHC. PHC is a complex process, but often underestimated, which requires the use of various diagnostic and imaging technologies (see Figure 1) in order to be successful. Its true value may not be realised if part of the tasks (identifying the concerned marker) remains unfinished
Men are only so good as their technical developments allow them to be.
– George Orwell
Discovering the marker that has the potential to stratify patients is only half the mystery—the way to measure markers that can be implemented into routine clinical practice, is the real challenge. The current discovery and validation of biomarkers is still based on an outdated research model that assumes a given treatment will react on a disease regardless of individual variability. Because of this approach, the burden of this breakthrough technology is placed in the later stage of drug development, rather than synchronised detection during pre-clinical and early stage drug development, with an emphasis on validation during the later phases. This alternative methodology could allow us to make considerably better decisions in late research and early development—essentially an evolution of the current discovery paradigm.
To reap the benefits of biomarker discovery in trial design, validated surrogate markers need to be identified to run shorter trials whereby response is measured based on the changes of these markers; and also smaller trials should be the aim, where potential responders can be determined through the detection and quantification of “response” markers. However, the rapidly growing understanding of the molecular basis of disease pathology, aided by progress in genomics, genetics and basic cell biology provides us with new insights into inherited differences among patients and disease states. This understanding enables the development of new diagnostics and more targeted medicines that are more effective and safer to be developed—the ultimate goal of “personalised healthcare”.

Diagnostics plays a decisive role in personalised healthcare. Research-use assays are able to quantify biomarkers to support decision making during drug development and to better understand disease pathways and mechanisms. Further, diagnostic tests would allow identification of patients who are most likely to respond to a specific treatment, helping physicians better diagnose and manage diseases at a patient level.
The development of any diagnostic technology involves challenges like identifying the right biomarker early enough, developing pharmacodiagnostics within drug timelines and ensuring sufficient collection of the right samples types, which bring into play countless regulatory and operational issues that can not be simply overcome when working across the globe. But, as Albert Einstein once said, “In the middle of difficulty lies great opportunity.”
The cost advantage of biomarkers in routine practice
The value of biomarkers is rarely in question, for example when one looks at the efficacy of widely used drugs of selected classes: ACE inhibitors (10-30%), beta blockers (15-35%), SSRIs (10-25%), TCA (20-50%) and statins (10-60%), a personalised concept is warranted. The value and cost of treatment is put into context when you consider if 65 previously well middle-aged men with high LDL, will take over 200,000 prescribed statin pills over the course of five years, when the likelihood that only one of those men would develop a stroke or transient ischemic attack without treatment. It’s probable that a biomarker measure could address the question of whether or not every patient should take a statin for high LDL (Low-density lipoprotein) levels. However, when the benefit risk ratio is evaluated for such an indication it may no longer warrant such a concentrated effort when the incidence of cardiovascular disease has dropped considerably in the last decades. There is also value with regards to costs and patient safety, with a recent meta-analysis of the incidence of serious (6.7%) and fatal (0.3%) ADRs in hospitalised patients claiming that 100,000 Americans die each year from drugs and 2.2 million Americans experience serious ADRs that act as an economic burden to the healthcare system.
There are obvious financial benefits of Personalised Medicines from the point of view of ever soaring healthcare costs, as the non-responders or poor responders are removed from the pool of users, the costs (both monetary and negative utility) for adverse events are avoided. Moreover, more precise targeting can lead to a greater volume of adoption by good responders who tend to have improved compliance and therefore additional net benefits, especially for long-term chronic therapies. Importantly, the ability to predict improvements or outcomes creates additional value for patients as they face less uncertainty when confronted with their disease prognosis.
Realising the true value
Biomarker discovery and validation is essentially the driver for many PHC treatments, however the return on investment when there is no guarantee of success, puts into question the “economic value” of such an approach. There is no real measure of the value of a new diagnostic test or targeted therapy but could be based on factors such as cost savings for both governments and patients, years of life gained, improvements in quality of life or morbidity and the decreased doubt on the potential outcome. There remains a fundamental question on how fast the progress of the science behind biomarkers can be translated into useful applications in drug development and clinical use. To help answer the scientific question of biomarkers, it is important for regulatory systems to be able to adopt a flexible approach and see the long-term benefits of patient related goals as well as cost savings. There is a need to encourage the basic science and information sharing for hypothesis generation in the clinical trial setting, to enable the personalised approach when assessing the overarching value of new treatments taking into account the balance of the appropriate patient safety considerations.
References
1. T. Andrew Clayton et al.: Pharmaco-metabonomic phenotyping and personalized drug treatment. Nature, Vol 440, 1073 -1077, 2006
2. Norton, Ronald. "Pharmacogenomics and Individualized Drug Therapy." Medscape Pharmacotherapy. 2001
3. Norton, Ronald. "Pharmacogenomics and Individualized Drug Therapy." Medscape Pharmacotherapy. 2001
4. Biomarkers Definitions Working Group.: Biomarkers and surrogate endpoints: preferred definitions and conceptual framework, Clin Pharmacol Ther. 69:89-95, 2001
5. National Diabetes Fact Sheet 2002, at http://www.cdc.gov/diabetes/pubs/estimates.htm#incidence (July 6, 2005).
6. U.S. Department of Health and Human Services, Public Health Service, Progress Review, Heart Disease and Stroke, April 23, 2003, at http://www.healthypeople.gov/data/2010prog/focus12/ (July 6, 2005).
7. "Roche Licenses Affymetrix’ GeneChip Technology." [Roche Press Release.] January 20, 2003. http://us.diagnostics.roche.com/press_room/2003/013003.htm
8. Lynch TJ et al.: Activating Mutations in the epidermal growth factor receptor underlying responsiveness of non-small cell lung cancer to Gefitinib. N Engl J Med, 350: 2129 - 2139, 2004
9. Carlini LE et al.: UGT1A7 and UGT1A9 polymorphisms predict response and toxicity in colorectal cancer patients treated with capecitabine/irinotecan. Clin Cancer Res.;11:1226-36, 2005.
10. Wen MS et al.: Prospective study of warfarin dosage requirements based on CYP2C9 and VKORC1 genotypes. Clin Pharmacol Ther. Jul 84:83-9, 2008
11. Flockhart DA et al.: Pharmacogenetic testing of CYP2C9 and VKORC1 alleles for warfarin. Genet Med. 10:139-50, 2008
12. van Kuilenburg AB, De Abreu RA, van Gennip AH: Pharmacogenetic and clinical aspects of dihydropyrimidine dehydrogenase deficiency. Ann Clin Biochem. Jan 40:41-5, 2003. Review
13. Spear et al, Trends in Molec Medicine 2001;7:201-204
14. Pederson TR, Lancet 1994;334:1383-1389 (Scandinavian Simvastatin Survival Study)
15. Lazarou et al, JAMA 1998;279:1200-1205, Spear et al, Trends in Molec Medicine 2001;7:201-204













