Impact of AI (LLMs & SLMs) in Clinical Trials for Precision Medicine
Kuldeep Jiwani, VP, Head of AI Solutions, ConcertAI
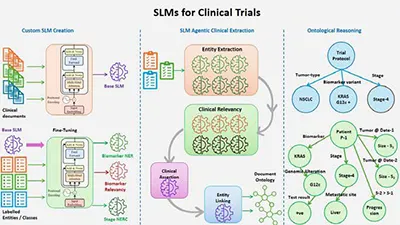
Precision Medicine demands accurate extraction of biomarker molecular data from clinical notes for targeted therapies. AI-driven custom-tuned SLMs, operating in a multi-agentic approach integrated with Medical Ontologies and Knowledge Graphs, enhance precision. This approach, spanning extraction, relevance, and reasoning, ensures high accuracy, zero hallucinations, and cost-effective processing of vast clinical data.

1. What are the key differences between general-purpose LLMs and custom-trained SLMs when applied to clinical entity extraction in precision medicine?
In the realm of healthcare and oncology, data sources primarily fall into two categories. The first consists of patient-related data, including structured information from electronic health records (EHRs), clinical notes, genomic reports, and similar records. The second encompasses publicly available medical literature, such as research journals and scholarly articles.
General-purpose Large Language Models (LLMs) like LLaMA and OpenAI models are trained on publicly accessible data. By design, they possess strong reasoning capabilities, enabling them to infer complex information with minimal instruction. Leveraging their understanding of medical literature, they perform reasonably well in extracting clinical entities from patient notes. For instance, identifying a nuanced concept like tumor progression status from clinical notes is a task that LLMs handle with commendable accuracy. However, in the field of precision medicine, "good" is not sufficient data extraction must achieve near-human levels of precision and recall while maintaining consistency. This presents a significant challenge for most general-purpose LLMs, as they struggle to sustain high precision (>0.9 F1 scores) due to variability in responses across different text patterns and prompts. Furthermore, the occurrence of ad-hoc hallucinations further complicates their reliability.
The root of this issue lies in the nature of publicly available medical resources. Medical blogs and research journals tend to be either highly technical or broadly informational, whereas physician-generated patient notes are distinct. These notes blend layman’s terms with specialised medical terminology, reflecting a patient’s unique medical journey. Consequently, the linguistic structure of patient records differs significantly from that of public medical texts.
This is where custom-trained small language models (SLMs), specifically tuned for patient notes, gain a crucial advantage. By learning the exact linguistic patterns found in clinical documentation, these models achieve superior precision and recall with greater consistency, making them a more reliable choice for precision medicine applications.
2. How do hallucinations in LLMs impact the accuracy of clinical data extraction, and what strategies can be employed to mitigate these risks in a high-stakes domain like precision medicine?
Hallucinations remain one of the most persistent challenges faced by LLMs. Even the latest advancements have not fully resolved this issue OpenAI’s most recent GPT-4.5 still exhibits a hallucination rate of 37.1 per cent, while its predecessor, GPT-4o, hallucinated at an even higher rate of 61.8 per cent. The same holds true for open-source models like LLaMA, which continue to struggle with generating consistently accurate responses.
This unpredictability poses a serious challenge, as it undermines the reliability of these models. One cannot expect consistent behavior from them, every third prediction may deviate from the expected pattern. For instance, consider a scenario where a carefully crafted prompt is designed to extract biomarker attributes, explicitly differentiating molecular variant types from genomic alterations. While the model might perform well on a limited set of test cases, its accuracy diminishes when applied to a broader range of documents. It may conflate the two categories or erroneously associate them with unrelated attributes. In high-stakes domains like precision medicine, such inconsistencies can have dire consequences.
The root of this issue lies in the way general-purpose LLMs operate. These models function based on instructions, generating an on-the-fly interpretation of data with every prompt. Since this process is inherently dynamic, even minor changes, such as altering a comma to a semicolon can shift the model’s behavior. Furthermore, complex or ambiguous clinical terminology can easily mislead the model into generating incorrect or fabricated information. Compounding this problem is the fact that these models are trained primarily on publicly available documents, which lack the depth required to fully grasp the intricate nuances of clinical contexts.
To illustrate this challenge through an analogy: when faced with an unknown illness, one initially consults a general physician. However, if the condition is severe or pertains to a specific organ, seeking a specialist becomes essential. Similarly, while general-purpose LLMs are effective for a wide range of tasks, they fall short in more critical applications. In such cases, specialized SLMs serve as the "experts" of the AI world. By being meticulously trained on domain-specific data, they significantly reduce the occurrence of hallucinations, making them far more reliable for precision-driven fields like precision medicine.
3. How do LLMs and SLMs compare in their ability to detect and interpret context-dependent clinical nuances, such as differentiating between similar genetic variations or treatment pathways?
In the field of clinical entity extraction for precision medicine, every subtle detail must be captured with utmost accuracy. Even the smallest discrepancy can lead to different targeted therapies, as there is a strong correlation between genetic variations and treatment pathways. For instance, two patients may both exhibit an EGFR-positive mutation, yet one may have a molecular variant classified as p.T790M, while the other presents with an Exon 20 insertion mutation. According to FDA-approved treatment protocols, the first patient could be prescribed Osimertinib, whereas the second might be treated with Amivantamab or Mobocertinib.
As previously discussed, LLMs are prone to hallucinations and often lack the specialized domain knowledge required to accurately capture genetic variations. This poses a significant challenge for research centers conducting clinical trials, where precise identification of specific molecular variations is crucial for recruiting eligible patients. To ensure reliable information extraction, models must achieve high F1 scores, balancing precision (to maintain accuracy) and recall (to ensure broad patient coverage for trials).
These critical variations are typically documented in physicians' clinical notes and genomic reports. If a sufficient volume of such data is available, a specialised SLM can be trained exclusively for this purpose. By focusing on a specific attribute and deeply understanding the nuances of clinical data and medical terminology having been trained on a domain-specific corpus such models can achieve superior performance. Through meticulous sampling strategies and fine-tuning techniques, precision and recall can be optimised, making SLMs far more effective in extracting clinically relevant information with both accuracy and consistency.
4. Considering the computational and financial costs of deploying LLMs for clinical trials, how do SLMs provide a more cost-effective alternative without compromising accuracy?
As previously discussed, SLMs can be fine-tuned to extract specific clinical data elements by training on real-world clinical notes. This makes them particularly effective for precision medicine, where accuracy in extracting clinical variables is crucial. For most key clinical attributes, SLMs outperform LLMs in precision and recall. However, for complex entities that require advanced reasoning, such as tumor progression status or metastatic status, SLMs alone may not suffice. In such cases, a chain of specialised SLMs, each dedicated to a specific subtask, can be employed. This agentic system ensures high accuracy even for intricate clinical extractions by leveraging a structured, multi-step approach.
Beyond accuracy, computational cost is another critical factor when comparing LLMs and SLMs. These models exist in various sizes, ranging from ultra-large models exceeding 100 billion parameters (e.g., 70B, 140B, or 405B) to more moderate LLMs in the 2B–20B range. The definition of an SLM varies, with some considering models as large as 8B parameters to fall within this category. However, for this discussion, we define SLMs as those ranging between 0.1B and 2B parameters.
In the context of clinical trials, where millions of patient records must be processed, ultra-large models are prohibitively expensive. Consider a comparison between an 8B LLM and a 0.4B SLM. A single patient may have 100 to 500 documents, with each document taking approximately 2 to 5 seconds to process on an 8B LLM, whereas a 0.4B SLM completes the same task in just 100 to 200 milliseconds. Furthermore, the LLM requires a high-end p4d.24xlarge AWS EC2 instance, while the SLM achieves comparable results on a significantly cheaper p3.2xlarge EC2 instance. Combining both factors, the 0.4B SLM is 25 times faster than the 8B LLM while running on hardware that is 10 times cheaper. This translates to an overall cost efficiency of approximately 250x, making SLMs the clear choice for large-scale clinical applications.
Another perspective on cost efficiency lies in the operational model of LLMs versus SLMs. To illustrate this, consider an analogy: imagine a factory that builds custom bicycles on demand. If employees receive a new, personalised bicycle every morning for their commute and return it at the end of the day, the factory must continuously manufacture new bicycles, an expensive and inefficient process. This is akin to how LLMs operate: each prompt dynamically generates a model instance on the fly, consuming significant computational resources. Now, suppose employees were given one high-quality bicycle, tailored to their needs, which they could use indefinitely. This would be far more cost-effective. Similarly, SLMs can be “frozen” for specific tasks, meaning once trained, they can consistently process documents without reconfiguring their instructions each time. This fundamental difference in execution makes SLMs not only faster and more precise but also significantly more economical for real-world clinical applications.
5. How can AI-powered information extraction enhance patient recruitment for clinical trials targeting rare genetic mutations, and what challenges still remain in ensuring high recall and precision?
The success of a clinical trial depends heavily on enrolling a sufficient number of eligible patients in accordance with the study protocol. With advancements in precision medicine, trial protocols are becoming increasingly specific, often targeting rare genetic mutations. Since these mutations occur infrequently, identifying suitable patients poses a significant challenge. For manual data curation teams, this task becomes monumental, if not impossible, when faced with the need to scan through hundreds of thousands of patient records in a short period. This is where AI-driven patient screening becomes indispensable, as machines can efficiently process vast volumes of data in a fraction of the time required by human teams.
However, an AI-powered machine curation system must overcome two primary challenges: accuracy and cost. Recent advancements in AI, particularly through LLMs and SLMs, have significantly improved accuracy across a broad range of clinical tasks. Moreover, when the task is well-defined, such as clinical entity extraction for a specific set of elements, cost efficiency becomes a crucial factor. In such cases, tailor-made SLMs offer a more cost-effective alternative to LLMs, delivering precise results while significantly reducing computational overhead.
By leveraging AI-driven automation, clinical trials can accelerate patient identification, improve screening efficiency, and enhance the feasibility of studies focusing on rare genetic mutations, ultimately driving medical innovation forward.
6. What role do domain-specific knowledge graphs and medical ontologies play in improving the reliability of AI-driven clinical data extraction?
There are two primary ways to leverage medical ontologies and knowledge graphs in clinical data extraction: advanced reasoning and standardising data into canonical forms.
In precision medicine, the quality of clinical trial data must be of the highest standard to ensure both accuracy and usability. However, the unstructured nature of clinical data poses a significant challenge. AI-driven systems extract information from clinical notes and reports, which are recorded by multiple physicians, each with their own way of expressing medical concepts. This linguistic variability creates a fundamental obstacle—how to reconcile diverse terminologies into a standardised framework.
To address this, AI systems rely on medical ontologies, many of which fall under the unified medical language system (UMLS) umbrella, alongside other standardised frameworks. By building semantic queries on these ontologies, AI can map multiple variations of a clinical concept to a standardised representation, complete with a corresponding code and vocabulary.
Medical ontologies also provide semantic relationships that help disambiguate complex concepts. For instance, consider the terms "Carcinoma of the left lung" and "Carcinoma of the right lung." While these are distinct diagnoses, they share a common parent category within the ontology: "Carcinoma of the lung." Depending on the level of granularity required, one can either drill down into specific details or zoom out for a broader representation of key concepts.
Beyond standard ontologies, patient-level document ontologies can be created to map a patient’s clinical journey by linking clinical notes and reports in chronological order. This enables a temporal representation of disease progression, such as tracking the evolution of a patient’s cancer stage over time. Moreover, knowledge graphs can be used to establish meaningful relationships between different clinical entities. For example, in a stage IV lung cancer patient, the primary tumor in the lung may have metastasised to the brain. This relationship can be structured in a document ontology, where nodes connect tumor stage, primary cancer site, and metastasis locations.
Such structured representations are invaluable in deriving complex clinical insights, such as tumor progression and disease trajectory, enabling more sophisticated and precise AI-driven clinical analysis.
7. What future advancements in AI and NLP do you foresee having the most transformative impact on clinical trial design and execution for precision medicine?
One of the most remarkable achievements of AI in recent years has been the advent of Large Language Models (LLMs), which have introduced several groundbreaking capabilities previously unseen. These models possess the ability to interpret instructions, engage in reasoning, and formulate responses in a structured, step-by-step manner—akin to human thought processes.
LLMs have excelled in domains where publicly available information is abundant. However, their effectiveness diminishes when applied to specialized fields that require domain-specific expertise. To bridge this gap,g retrieval-augmented generation (RAG) systems were introduced. These systems dynamically retrieve relevant domain knowledge and feed it into LLMs, enabling them to make more informed decisions. When combined with knowledge graphs, this approach proved effective in expanding AI’s capabilities in certain specialized areas. Yet, even these enhancements reached a natural limit, unable to fully compensate for the models’ lack of inherent domain expertise.
An alternative solution involved pretraining LLMs from scratch on specialised domains. While this significantly improved performance, the process proved to be prohibitively expensive and resource-intensive. In response, SLMs emerged as a practical middle ground, offering a balance between effectiveness and computational efficiency. However, to achieve broader objectives, multiple SLMs often need to work in tandem, adding complexity to their deployment.
The next major breakthrough in AI will likely come when models evolve beyond mere text-based reasoning and begin to mimic human knowledge abstraction more closely. This advancement would allow AI to understand and interpret concepts contextually, much like the human mind does.
For instance, when encountering the term EGFR, a human immediately considers it as the epidermal growth factor receptor biomarker. However, if the term is followed by "mL/min," the context shifts—EGFR now refers to the estimated glomerular filtration rate, a measure of kidney function. Further, if the phrase appears within a descriptive report rather than a patient’s test result, its relevance and interpretation change once again. This ability to dynamically contextualise information is what current AI models lack but must achieve for true reasoning capabilities.
Since human cognition operates through interconnected subjects and objects, knowledge can be systematically represented using subject-predicate-object (SPO) triplets in a graphbased framework. If AI models are trained on a universal, large-scale knowledge graph, the next generation of Language Concept Models could autonomously extract, abstract, and interpret complex information.
Such a breakthrough would have profound implications for clinical trials and precision medicine. Machines would gain the ability to identify intricate medical patterns at an unprecedented scale, revolutionising drug development and personalized treatment strategies. By enhancing AI’s ability to comprehend and generalise complex medical phenomena, research institutions could accelerate the discovery of more effective therapies, ushering in a new era of medical innovation.











