How to Use Pharmacogenomic Information to Reduce the Risk of Experiencing Adverse Events
Giovanni Furlan, Fellow, International Society of Pharmacovigilance
Genetic testing can be used to reduce the risk of experiencing adverse reactions. Seldom is one single gene responsible for altering a drug’s benefitrisk balance. This is more often due to multiple genetic and non-genetic factors, thus increasing the complexity and limiting the utility of a widespread use of pharmacogenomics. Widespread use of genotyping and artificial intelligence can overcome these limits.

Pharmacogenomics is the study of the genes, their structure, function, polymorphisms, transcription, and translation, how they interact with each other and with environment. The main goal of pharmacogenomics is to understand how genetic variants can alter a drug’s benefit-risk balance. Genetic differences that are likely to be of most relevance to drugs and that can alter their benefit-risk balance pertain to four broad categories:
• Genes that affect proteins involved in the metabolism of a drug, its active metabolites, or their transport. Variants affecting these genes are mainly associated with pharmacokinetic alterations and might require a drug dose adjustment
• Genes that code for proteins that are intended or unintended drug targets and other pathways related to a drug’s pharmacological effect. Variants of these genes can shift the relationship between the drug dose and its effect, thus requiring a change in the administered drug dose
• Genes not directly involved in the drug’s pharmacology that can alter an “off-target” protein. In this instance idiosyncratic reactions (e.g., immune mediated adverse reactions) can occur
• Genes that influence disease susceptibility or progression. Variants of these genes can become therapeutic targets.
These genetic variants are estimated to account for 15 per cent-20 per cent of the different desirable and undesirable responses that patients have following drug intake and for some drugs they can account for up to 95 per cent of interindividual variability.
Multiple genetic and non-genetic factors can increase the risk of experiencing an adverse reaction: the example of warfarin
It is rare that one single gene variant is responsible for altering the metabolism of a drug or for causing an idiosyncratic adverse reaction. More often, as in the case of warfarin, (a drug used for the prevention of thrombosis and thromboembolism) multiple genetic and environmental factors are involved. This drug is of particular interest since it has a narrow therapeutic index, the inter-individual variability of the dose required to achieve the required therapeutic effect can vary up to 20 times and an excessive dose increases the risk of bleeding, while a subtherapeutic dose increases the risk thromboembolism.
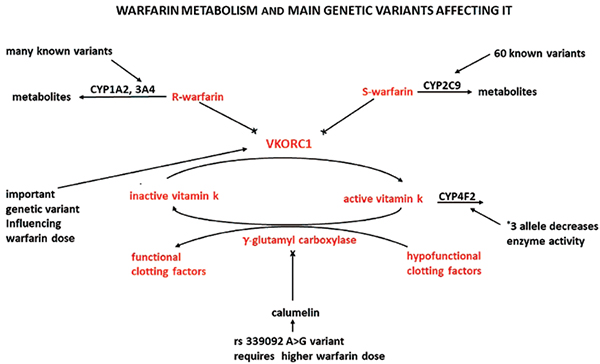
To identifywhich genetic variants are of importance, boththe metabolic steps and the mechanism of action of warfarin need to be considered.
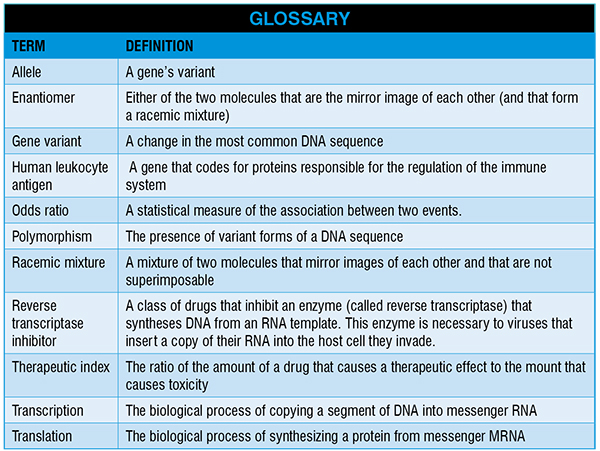
As shown in figure 1, warfarin is a racemic mixture, and the S-enantiomer is more potent in inhibiting vitamin K epoxide reductase (VKORC1) as compared to the R-enantiomer. (Note: VKORC1 reduces vitamin K, an essential cofactor of gamma-glutamylcarboxylase, the enzyme that converts hypofunctional clotting factors to functional). It is, therefore, not surprising that genetic variants affecting cytochrome 2C9 (CYP2C9), the enzyme that metabolises the S-enantiomer, have a greater influence in altering the dose of warfarin that is needed to achieve the therapeutic effect as compared to genetic variants of CYP1A2 and 3A4, since these cytochromes are responsible for the metabolism of the less potentR-enantiomer. The variability in the needed warfarin dose is partially explained by the more than 60 known variants affecting CYP2C9 and, even if not all these variants are important, their prevalence varies from one ethnicity to another. CYP2C9 *2 variant, for example, requires a lower warfarin dose, but its frequency is around 13-14 per cent in Caucasians and less than 1 per cent in Asian patients.CYP2C9*5,*6,*8 and *11 polymorphisms also require a lower warfarin dose, but their prevalence varies from 10 per cent in Africans to less than 1 per cent in Caucasians. There is another variant in a non-coding region of CYP2C9DNA that requires to reduce warfarin dose only in African Americans. The reason is not clear, but it could be that only in this population this variant is inherited together with another polymorphism that affects cytochrome metabolism.
Another very important genetic polymorphism reduces by twofold the expression of VKORC1 and, therefore, requires a lower dose of warfarin. The frequency of this allele is around 90 per cent in Asiatic patients, while it is only round 10 per cent in African Americans. Also, the gene coding for CYP4F2, that metabolises reduced vitamin K to hydroxy vitamin K, has a variant(CYP4F2*3) that lowers the concentration of this cytochrome thus increasing active vitamin K concentration and requiring a higher warfarin dose. As for the previously detailed variants, also this allele has a different prevalence in different ethnicities: it is presence ranges from 24 per cent in Caucasians to 7 per cent in African Americans.
Finally, calumelin, a protein that inhibits gamma-glutamyl carboxylase, is characterised by a variant that requires African Americans to take a lower warfarin dose to achieve the targeted therapeutic effect.
The multitude of variants that influence the required warfarin dose and whose prevalence varies in the different ethnicities, explains why traditional dosing algorithms, that only look at CYP2C9 *2 and *3 together with the above mentioned VKORC variant, might not be optimal for all patients regardless of their ethnicity. Algorithms should rather be adapted to consider which variants can influence the required drug dose in the various ethnicities. However, more precise dosing algorithms should also consider non-genetic factors. In fact, in the case of warfarin also elderly age, high weight, amount of vitamin K taken with the diet, drugdrug interactions involving the drug and tobacco smoking alter the required dose.
Factors that influence the need for genetic testing
Since many genetic variants can influence the effect of a drug and it would be unpractical and expensive to perform genetic testing for all the involved alleles, the logical question is when genetic testing should be performed and for which variants. To answer this question, the strength of evidence associating an allele to an adverse event and the expected consequences of the adverse reaction need to be determined. Therefore, it is necessary to evaluatethe severity and seriousness of the adverse event associated with the genetic variants, since, for example, it is not worthwhile to investigate the contribution of a genetic polymorphisms to a mild localised erythema associated with a drug used for cancer. Other aspects that have to be considered are the magnitude of the adverse event increased frequency, the probability with which a genetic test will indicate the patients have the genetic variant of interest among those who actually have the variant (i.e., the analytical sensitivity), the accuracy with which a genetic test predicts the clinical disorder (i.e. the clinical validity of testing)and the different frequencies in the various ethnicities of all the genetic variants contributing to an adverse event.All these factors contribute to the genetic test positive predictive value that tells us what is the probability that a patient taking the drug of interest and with a positive test for a specific genetic allele will experience the adverse event.
The more genetic and non-genetic variants contribute to an adverse event, the less it is likely the genetic variant of interest will cause an adverse event: therefore, its positive predictive value will be low.
Examples of genetic testing utility
Not all adverse reactions are characterised by a complex interplay of multiple genetic and non-genetic factors as is the case of warfarin.
Abacavir, is a reverse transcriptase inhibitor indicated for human immunodeficiency virus infection (HIV). It can cause a hypersensitivity syndrome characterised by respiratory, gastrointestinal disorders and rash. In patients who are rechallenged with abacavir after having experienced a hypersensitivity syndrome, this adverse reaction can be fatal. A study in about 1900 patients showed that avoiding abacavir in patients with human leukocyte antigen (HLA)-B*57:01 variant eliminated immunologically confirmed hypersensitivity reactions. In this study, the test negative predictive value was100 per cent: this meant all the patients that had a negative genetic test did not experience the adverse reaction. Considering an overall prevalence of HLA-B*57:01 variant of 6 per cent and being the positive predictive value of the test of 47.9 per cent,it has been calculated that 25 patients need to be screened to avoid one case of abacavir induced hypersensitivity syndrome.Therefore, the medicine’s Summary of Product Characteristics requires that before taking abacavir all patients are screened for HLA-B*57:01 variant; patients who are found to carry this allele should not be treated with abacavir.
A very different example is that of flucloxacillin, a medicine used to treat bacterial infections. The strongest known genetic association for drug-induced liver injury is that between flucloxacillin and HLA-B*57:01 (the odds ratio of the association is 80). The genetic test accurately identifies the patients who have the variant, but flucloxacillin drug induced liver injury occurs (luckily) in only 8.5 out if 100,000 patients who take the drug and the positive predictive value of the test for the HLA-*B57:01 allele is of 0.12 per cent. It has been calculated that around 13,500 patients need to be screened to avoid one case of drug induced liver injury and due to the low positive predictive value of the test, only 1 out of about 830 patients who take flucloxacillin and have the HLA-B*57:01 variant will suffer from drug-induced liver injury.Considering that the prevalence of the HLA-B*57:01 variant is relatively high (in Caucasians, for example, it is of around 7 per cent) and that only a small fraction of patients with the variant will experience drug-induced liver injury due to flucloxacillin, it has to be decided if these patients should take the medicine or if they should be denied flucloxacillin. Another non-trivial point to examine is the cost-effectiveness of screening around 13,500 patients to avoid one case of liver injury.
Genetic testing clinical utility is therefore being pondered: what is the balance of benefits and risk associated with using a test in everyday practice, including its ability to inform clinical decision making and prevent adverse reactions.
As for abacavir, also for flucloxacillin, the answer can be found in the medicine’s Summary of Product Characteristics: HLA-B*57:01 variant is only mentioned in section 4.8 ‘Undesirable Effects’ where it is stated there is evidence that the risk of flucloxacillin induced liver injury is increased in patients who carry the abovementioned allele. However, since very few carriers of the allele will develop liver injury, routine screening for the variant is not recommended.
A look in the future of personalised medicines: pros and cons
The effect of a genetic variant on an adverse reaction varies from being the most important risk factor to being only a minor contributor. In the future, it is possible that all the gene variants affecting the efficacy and safety of a drug and how they interact with the environment will be known. The human population could all be genotyped, and the genotype could be included in electronic medical records so that artificial intelligence can consider all the individual variables and tell which is the best medicinal product and the most appropriate dose for each patient suffering from a pathology. Therefore, the benefit-risk balance of a drug could be assessed based on individual needs.
However, like everything pertaining to this world, there will also be some cons. For example, the data on the genetic characteristics of a person could be hacked and used to discriminate some individuals. Health insurance companies might require access to individuals’ genotype, and the cost of health insurance could be higher for subjects with genes predisposing to certain pathologies. Will everyone really want to know, through the analysis of our genotype, what is our risk of suffering from a pathology, or will this only increase people’s anxiety? Finally, it is uncertain if everyone will understand the meaning of their genetic characteristics, especially since in an era where infodemic and fake news are rife. Unluckily, it is easy to imagine that those who will not understand the meaning of their genetic characteristics will look for information in the web and will easily fall prey to those who want to sell improbable remedies for their risk factors or, even worse, who have political agendas.
References:
1) Pharmacogenomics. Challenges and Opportunities in Therapeutic implementation. Lam Y.W.F. & Scott S.A. Editors, Academic Press 2 nd Edition.
2) U.S. Department of Health and Human Services. Clinical Pharmacogenomics: Premarket Evaluation in Early-Phase clinical Studies and Recommendations for Labeling. Guidance for the Industry. January 2013
3) Hogg K &WietzJ.I. Blood Coagulation and Anticoagulant. Fibrinolytic, and Antiplatelet Drugs. In Goodman & Gilman’s The Pharmacological Basis of Therapeutics. Bruton, Hilal-Dandan and Knollman Editors. 13th Edition, Mc Graw Hill.
4) Cavallari L.H&MomaryK.M. Pharmacogenomics in Cardiovascular Disease. In: Lam Y.W& Scott S.A Editors, Pharmacogenomics. 2 nd Edition, Academic Press
5) Dean L. Warfarin Therapy and VKORC1 and CYP Genotype. 2012 (Updated in 2018). In: Pratt VE et Editors. Medical Genetic Summaries. Bethesda: National Center of Biotechnology Information.
6) Daly A.K. Pharmacogenomics of Warfarin. In Handbook of Pharmacogenomics and Stratified Medicine. 2014 Elsevier
7) Johnson JA et al. CPIC Guideline for Pharmacogenomics-Guided Warfarin Dosing: 2017 Update. Clinical Pharcol& Therap. 2017. 102 (3): 397-404
8) Burke W. Genetic Tests: Clinical Validity and Clinical Utility. CurrProtoc Hum Genet. 2014;81:9.15.1-9.15.8.
9) Mallal S et al. HLA-B*5701 Screening for Hypersensitivity to Abacavir. N Engl J Med 2008; 358:568-579
10) Turner R.M. &PirmohamedM. Pharmacogenomics of Adverse Drug Reactions. In Grech G, Grossman I Editors. Preventative and Predictive Genetics: Towards Personalised Medicine. 2015 Springer
11) ViiV Healthcare Uk Ltd. Ziagen 300 mg film-coated tablets Summary of Product Characteristics. Last updated 05 October 2021. Accessed on 25 May 2022 at:https://www.medicines.org.uk/emc/product/5518/smpc#gref
12) Alfirevic A &Pirmohamed M. Predictive genetic testing for drug-induced liver injury: considerations of clinical utility. Clin PharmacolTher. 2012 Sep;92(3):376-80
13) Aurobindo Pharma. Flucloxacillin 500 mg capsules Summary of Product Characteristics. Last updated on 25 January 2021. Accessed on 25 May 2022 at:https://www.medicines.org.uk/emc/product/545/smpc#gref













