Target Deconvolution in the Post-genomic Era
Georg C Terstappen, Chief Scientific Officer, Siena Biotech S.p.A., Italy
In times of reduced productivity coupled with increasing costs within the pharmaceutical industry, reductionistic target-based drug discovery has come under pressure and phenotype-based discovery has gained momentum. Target deconvolution is an important aspect for efficient progression of compounds towards development.
While the growing demand for innovative medicines is creating increasing opportunities for the pharmaceutical industry, it is exactly this industry that suffers from an expensive disease itself, known by the name "innovation deficit". In fact, while the FDA approved 53 New Molecular Entities (NMEs) and biologics in 1996, a nearly steady decline since then has led to a disappointing 22 NME approvals in 2006, which is even more disappointing owing to the fact that twice as much money was spent. The cause of this malaise is of multiple nature and many different legal, political and financial factors have increased the complexity of current pharmaceutical R&D.
Most compounds-especially those with a new mechanism of action-fail due to a lack of efficacy observed during clinical development. This is mainly a consequence of insufficient understanding of disease pathophysiology in conjunction with a "reductionistic" target-based approach for drug discovery. In the 1990s, this approach largely replaced the more holistic approach of screening compounds for phenotypic changes in complex systems such as living animals and isolated organs (Figure 1). The target-based approach was thought to be more rational and efficient, largely corroborated by the sequencing of the human genome which was in progress at that time and promised the advent of a new era with all potential drug targets known.
Renaissance of phenotype-based drug discovery
The perceived "failure" of target-based drug discovery in current post-genomic times has led to a renaissance of the more holistic approach which allows identification of compounds with multiple cellular targets ("polypharmacologic compounds") since phenotypic effects are measured as an integral of the response of an entire system, thus better matching the polygenic and / or multifactorial nature of most diseases. Although many model systems such as worm, fly, zebrafish and mouse can be employed for small-scale drug screening, at present mainly mammalian cells in culture are compatible with High Throughput Screening (HTS) with a phenotypic read-out. Permanent cell lines are easy to handle, can be grown in large quantities and many are commercially available as well, and therefore are used predominantly. Furthermore, through recombinant gene technology aspects of disease biology can be engineered into these cells.

Figure 1
In the research laboratories of Siena Biotech S.p.A, dedicated to the study of Huntington's disease-a progressive neurodegenerative disorder-a stable human HEK293 cell line was developed which expresses the mutated huntingtin gene under the control of an inducible promoter. The recombinant cell system also contains a luciferase reporter gene under control of a cAMP-Responsive Element (CRE) which allows measurement of transcriptional activation. If the mutated huntingtin, which causes the disease in humans, is switched on, cells start to degenerate leading to transcriptional dysregulation and cell death. While the first is measured with the reporter gene, the latter phenotypic effect is simply determined by using a toxicity assay. A HTS with this recombinant cellular disease system has led to the identification of drug-like organic molecules that rescue the phenotypic effects caused by expression of the mutated huntingtin.
Target deconvolution
Whatever the disease model system that might have been used, the retrospective identification of the molecular targets underlying the observed phenotypic responses, termed target deconvolution, is important. From a basic research perspective, knowledge of the molecular targets will help to understand the (patho) mechanism(s) of the disease. But this knowledge will also aid rational drug design and allow efficient Structure-Activity Relationship (SAR) studies in a chemical optimisation programme, thereby developing target-specific assays. In addition, the aspects related to target-specific toxicity and side effects can be addressed, reducing later-stage "attrition" early on.
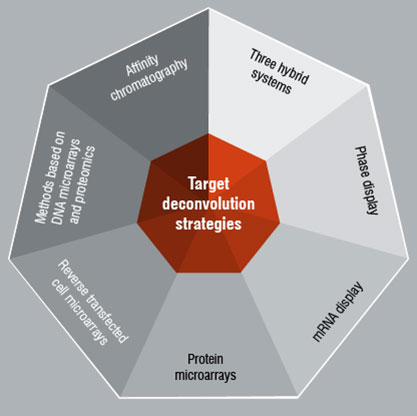
A wide range of different technologies can be employed for target deconvolution (Figure 2). However, no "best practice" or "standard operating procedure" is available and the method of choice will have to be decided on a case-by-case basis often mainly governed by the properties of the "drug" molecule. Methods leading to direct identification of targets typically exploit the affinity between the "drug" molecule and its target protein assuming that physical interaction is a prerequisite for functional effects.
Direct target deconvolution methods
"Classical" affinity chromatography-based methods have been successful for target deconvolution. An important advantage in these methods is that target proteins maintain their 3-D structure including post-translational modifications. Protein microarray-based strategies represent an alternative to affinity chromatography and identification of proteins which are expressed at low levels in vivo is not a priori compromised since large numbers of individually purified proteins are arrayed in equal amounts. Clearly, only those proteins can be identified that have been immobilised on the microarray. A method referred to as "reverse-transfected" microarrays employs microarrays of cells expressing defined cDNAs thus circumventing the need to use large numbers of individually purified proteins.
Technologies based on expression cloning (e.g. three hybrid systems, phage display and mRNA display) also include an affinity step. Since proteins are expressed from fusion constructs of cDNA libraries, the properties of these recombinant proteins might be different from their native counterparts, especially in terms of post-translational modifications. In addition, sophisticated molecular biology skills are needed, which is not the case for affinity chromatography. Common to all these direct methods is that the "drug" molecule has to be modified chemically for labelling and / or immobilisation purposes. This means that access to synthetic chemistry might be necessary and often a limited structure-activity relationship study will also have to be carried out in order to identify a position where the molecule can be modified under retention of its biological activity, which has to be tested.
Methods supporting target deconvolution
Target deconvolution can be supported by technologies based on comprehensive DNA microarray and proteomics analyses. In a more indirect way, application of these technologies, which do not need modification of the chemical molecules, can eventually also lead to the identification of the molecular targets. For instance, investigation of global gene expression changes of an organism induced by the "drug" molecule and determined by DNA microarray analysis can help to understand the MoA of the molecule and in the form of "connectivity maps" can be used to connect chemical molecules, genes and disease in a systematic manner. Global changes of the protein constitution of an organism in response to exposure to a "drug" molecule, studied by proteomics technologies, allow the generation of a "proteomic compound signature", again aiding to understand the MoA of the chemical molecule.
Confirmation of 'deconvoluted' targets
The ultimate goal of all strategies and methods for target deconvolution is the identification of the biological molecules that directly interact with the "drug" molecule. In addition, it is also important to confirm that modulation of the identified biological target is associated with the functional effects detectable in the phenotypic assay system that was used for identification of the chemical molecule. If 3-D protein structures are available, a structural analysis of the protein-compound interaction can be used to evaluate the interaction profile further. Since these in silico analyses generate predictions, direct proof of "drug"-protein interaction and its functional consequences is necessary. A variety of methods can be used to confirm physical interaction. For instance, surface plasmon resonance, and associated phenotypic changes can be confirmed by functional studies employing genetic methods such as RNA interference, leading to a knock-down of the target gene or gene overexpression. In vivo, transgenic and knock-out animals can be generated for detailed studies of phenotype.
Conclusion
Around the world, target-based drug discovery, carried out by major pharmaceutical companies and the biotechnology industry, has so far resulted in disappointing results in terms of delivering efficacious new medicines for therapy of diseases. This perceived "failure", which is associated with tremendous costs, has recently led to the renaissance of phenotype-based drug discovery. For this approach, typically cell-based assay systems are used that mimic aspects of disease biology or host disease-relevant pathways. Employing these "unbiased" systems, small organic molecules are screened for relevant phenotypic effects which might be mediated by an active compound's interaction with one or more cellular targets. Identification of these targets is important in order to progress active compounds efficiently through the drug discovery process. Although a broad panel of experimental target deconvolution strategies exist, none of these methods could universally be applied and hence a case-by-case evaluation is necessary. Apart from the choice of the method another important decision will be when to start deconvolution activities during the progression of active chemical molecules since timelines might be long and outcomes unpredictable. In this context, it might be comforting to know that for some of the approved drugs on the market, as of today the molecular target(s) are still not known.












