Towards Precision Medicine for Cancer Pain Treatment in Asian Populations
Yow Hui-Yin, Pharmacy Lecturer, Department of Pharmaceutical Life Sciences, Faculty of Pharmacy, Universiti Malaya
Shobha Elizabeth Satkunananthan, Master student, School of Pharmacy, Taylor's University
Vijay Suppiah, Senior Lecturer in Pharmacy, Clinical & Health Sciences at the University of South Australia
Toh Gaik-Theng, Senior Lecturer, School of Medicine, Taylor's University
Cancer pain is one of the most debilitating symptoms and opioids are the cornerstone of therapy in its management. However, opioid treatment outcomes are unpredictable due to inter-individual variability. Single nucleotide polymorphisms in genes encoding drug-metabolising enzymes (CYP2D6, UGT2B7), neurotransmitter metabolising enzymes (COMT, ABAT), drug transporters (ABCB1) and drug targets, including receptors (OPRM1) and ion channels (P2RX7) have been reported to be associated to inter-individual variability of opioid responses among patients of Asian ethnicities.

The prevalence of cancer pain is high among patients with advanced cancer, affecting up to 96 per cent of patients, which significantly impacts their quality of life. However, inadequate pain relief remains a serious health concern for cancer patients, particularly in Asia, where the prevalence of undertreated cancer pain is relatively high. Strong opioids, such as morphine, have been the mainstay of treatment for cancer pain based on three-step analgesic ladder of the World Health Organization (WHO). However, inter-individual differences in analgesic response pose a major disadvantage for the clinical use of these opioids. Pharmacogenomics, the study of a person’s genetic make-up and its effect on treatment response, provide a reasonable explanation for the inter-individual variability observed in analgesic response, particularly in long-term opioid use. While genetic variants associated with opioid treatment outcomes in patients with cancer pain have been identified, there have been inconsistent findings between studies. This review aimed to summarise current evidence in the association of genetic variants with treatment responses in Asian patients treated with opioids for cancer pain.
Materials and methods
A literature search was conducted in Medline and Embase using free-text and MeSH terms grouped into four main subject groups: cancer pain, pharmacogenetic, opioids and treatment outcome. Only original research articles published in English from the year 2000 to November 2020 investigating the associations between single nucleotide polymorphisms (SNPs) and treatment outcomes in cancer pain patients treated with opioids for pain were included. A total of 658 records were screened and 337 articles were shortlisted after removing duplicates and those published before 2000. Furthermore, a full text review resulted in 14 articles that fulfilled the inclusion criteria. The included studies assessed pain control and opioid dosage as treatment outcomes, and findings were presented according to the opioids studied in included articles, namely morphine, fentanyl, oxycodone, tramadol, codeine, hydrocodone, hydromorphone, levorphanol, methadone, and oxymorphone.
Results
Seventeen SNPs from eight genes were significantly associated with opioid dosage, pharmacokinetics, and pain control among Asian populations (Table 1).
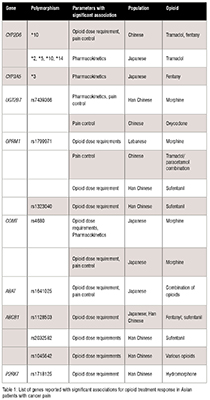
Discussion
This review focused on the impact of genetic variations on opioid responses in Asian patients with cancer pain. The study identified genetic variants in drug-metabolising enzymes (CYP2D6, CYP3A5, UGT2B7), neurotransmitter-metabolising enzymes (COMT, ABAT), drug transporters (ABCB1), and drug targets such as receptors (OPRM1) and ion channels (P2RX7) as contributing factors to the observed inter-individual variability in treatment outcomes.
Of these genetic variants, CYP2D6 polymorphisms were the most extensively studied, with the CYP2D6*10 allele being associated with higher tramadol dosage requirements and reduced analgesic effects due to decreased plasma concentration of metabolites. Additionally, the CYP2D6*10 variant was also associated with higher fentanyl dose and pain scores, despite not being involved in fentanyl’s metabolism. This CYP2D6*10 allele is present in up to 50 per cent of individuals of Asian ethnicities. The CYP3A5*3 allele was found to impact fentanyl’s pharmacokinetics as CYP3A4 and CYP3A5 are responsible for its metabolism. Further research investigating the potential relationship between CYP2D6*10 and fentanyl pharmacokinetics may explain the observed association.
Genetic variations in UGT2B7 affect morphine glucuronidation in forming its metabolites - morphine-3-glucuronide (M3G) and morphine-6 glucuronide (M6G). The rs7439366*C allele was observed to enhance analgesic efficacy and was significantly associated with higher levels of M3G and M6G. The T allele carriers were observed to have a reduced response to morphine and prolonged-release oxycodone. Additional studies are required to investigate this relationship as the level of evidence for this SNP is low (between levels 3 to 4 in PharmGKB) as this is currently insufficient to be used as a biomarker for drug responses.
Opioids bind to the μ-opioid receptor to bring about opioid-induced analgesia. The OPRM1 G472A (rs1799971) SNP, which is present at a high frequency in Asian populations, is associated with reduced analgesic efficacy of morphine requiring higher doses in affected individuals. The G allele of the SNP is thought to reduce signalling efficacy and expression of the μ-opioid receptor, leading to decreased binding potential resulting in increased morphine requirement. On the other hand, heterozygous patients for both rs1799971 and rs1323040 SNPs required a higher dose of sufentanil, suggesting of presence of a variant that reduced opioid analgesic efficacy.
The catechol-O-methyltransferase (COMT) enzyme is responsible for breaking down dopamine in the prefrontal cortex, and the COMT G472A (rs4680) SNP had been linked to various neurological conditions, including pain modulation and opioid-related disorders. Carriers of the AA genotype of this SNP were found to have lower plasma concentrations and dose requirements for morphine, while the carriers of the GG genotype had higher dose requirements. However, this finding was inconsistent with studies associating the A allele with reduced analgesic efficacy. The A allele carriers may have a higher sensitivity to pain but require less morphine doses. Pooled data from a meta-analysis found that the opioid consumption of patients who were COMT rs4680 AA and GG homozygotes was not significantly different in the first 24 hours of opioid treatment, while AG genotypes required significantly lower doses of opioids.
The rs1641025 SNP corresponds to a substitution from a T to C nucleotide within ABAT, which encodes GABA transaminase. Studies suggested that the ABAT rs1641025*C allele was associated with lower pain scores and enhanced opioid efficacy. While there is a lack of studies to show the clinical significance of this association, it is thought to modulate pain perception through the GABAergic synapse pathway by regulating physiological functions including pain perception. Further studies are needed to explore the potential pharmacogenetic role of this SNP in pain control and opioid analgesic responsiveness and possibly its usefulness as a biomarker.
The bioavailability of opioids is affected by P-glycoprotein (P-gp), a family of efflux transporters encoded by ABCB1. Three polymorphisms, namely rs1045642, rs2032582, and rs1128503 in ABCB1 have been extensively studied. Patients carrying the rs1045642*T allele have been found to have reduced expression of P-gp, potentially resulting in reduced efficacy or higher toxicity from opioids treatments. Studies on the three SNPs in patients receiving multiple opioids including fentanyl and sufentanil have produced mixed results, with some suggesting enhanced analgesic efficacy in carriers of the T allele while others showing reduced efficacy. This warrants further investigations to elucidate the impact of these SNPs on opioid effectiveness and toxicity.
The purinergic receptor P2X7 was associated with chronic inflammatory and neuropathic pain. In patients with lung cancer, the P2RX7 rs1718125 SNP was significantly associated with postoperative pain and fentanyl dose, with A allele homozygotes requiring the highest dose for pain control and having significantly higher pain scores than the GG genotype group. However, this finding contradicted a recent study that reported that carriers of the A allele required lower fentanyl and had better pain scores than those who were allele G homozygotes. The association of P2RX7 polymorphisms and opioid dose requirements and pain control needs further investigation, as preclinical studies suggested this gene may contribute to pain modulation both in peripheral tissue as well as processing in the central nervous system.
Most of the pharmacogenetic studies in Asian patients with cancer pain treated with opioids have focused on patients with East Asian ancestries, which may not be representative of other Asian populations. There were also differences in outcome measures used in these studies, making comparison of findings challenging. Compared to pharmacogenetic studies in Caucasian and European populations, the sample sizes in Asian populations tend to be smaller and focused on a limited number of genes and SNPs. Nevertheless, this review highlighted the findings on the association of genetic variants with treatment responses in Asian patients treated with opioids for cancer pain. The CYP2D6*10 SNP was found to be relevant to tramadol pharmacokinetics and opioid dose requirement. SNPs in UGT2B7, ABAT, and P2RX7 have demonstrated potential clinical relevance, while SNPs in OPRM1, COMT, and ABCB1 were found to have low clinical significance due to inconsistent findings. Several ABCB1 SNPs have been reported to be associated with opioid treatment outcomes, but further investigations are required to understand their role.
Conclusion
Variability in opioid responses was linked to genetic polymorphisms in drug-metabolising enzymes, neurotransmitter-metabolising enzymes, drug transporters, receptors, and ion channels. The clinical significance of OPRM1 rs1799971, COMT rs4680, and ABCB1 SNPs need further exploration in Asian patients with cancer pain.
Citation
Satkunananthan, S.E.; Suppiah, V.; Toh, G.-T.; Yow, H.-Y. Pharmacogenomics of Cancer Pain Treatment Outcomes in Asian Populations: A Review. J. Pers. Med. 2022, 12, 1927. https://doi.org/10.3390/jpm12111927
Funding
This research was funded by the Ministry of Higher Education (MOHE) Malaysia for Fundamental Research Grant Scheme (FRGS/1/2019/SKK09/TAYLOR/03/1).
Competing interests
The authors declare no conflicts of interest.















