There is significant concern that Personalised Medicine will cause the end of the blockbuster era in pharmaceuticals. The reality is that what must evolve is the model for financial return, application of stratification in medicine and focussed research on drug safety.
The pharmaceutical industry appears to be at a nexus of its evolution. It has been observed that new drug pipelines are less than adequate to continue its recent rate of growth and off-target safety issues are causing critical post-marketing failures. Added to this global economic pressures may impose greater constraints on drug pricing and the introduction of personalised medicine suggest a reduction in the marketing opportunity for new drugs based on genetic make-up of the patient. Collectively, these issues indicate that the concept of the blockbuster drug needs to evolve to address these challenges.
Redefining “Blockbuster”
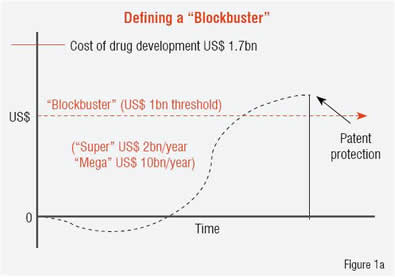
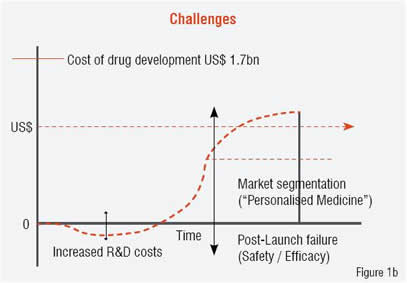
A “blockbuster drug” is conventionally defined as one that produces sales revenue in excess of US$ 1 billion per year, with “super-blockbusters” reaching US$ 2 billion per year. This barrier contrasts with the current cost-estimate for new drug development reaching US$ 1.7 billion. Currently there are approximately 100 drugs that meet the blockbuster definition with only five attaining super-blockbuster status (Lipitor, Plavix, Celebrex, Prilosec and Nexium). Figure 1 contrasts the existing blockbuster model (Figure 1a) with the challenges to that model that are noted above (Figure 1b). A significant opportunity exists to overcome these challenges by redefining “blockbuster” from annual revenue levels to the total revenue produced for the company while under patent protection (i.e. the integral, not the peak in Figure 2). Several changes in the drug development process would facilitate a transition to this new model, and personalised medicine, which is currently viewed as a threat to the existing markets, can provide a positive impact if implemented in a strategic manner.
Stratified medicine or personalised medicine
For some time, the spectre of personalised medicine (or pharmacogenomics as it frequently called) has met with concern in the pharmaceutical industry because of the impression that it would reduce the market potential for new drugs. This centres on the application of genomic screening of individuals to determine their potential response to a specific drug, e.g. presence of SNPs in a population or individual, and the potential impact on labelling a drug to address a significantly reduced population. As the technology continues to become more cost-effective, the collection of detailed polymorphisms in the population increases and the appropriate integration of the data into knowledge bases proceeds, regulatory agencies, e.g. FDA, EMEA, will further require such testing and incorporation into labelling guidelines for new drugs. The key question to address is: to what granularity can personalised medicine be pushed and still remain economically feasible for both drug development and the provision of healthcare while delivering the best quality of life for the patient? Most physicians would already contend that medicine is practiced in a “personalised manner” although pressures from reimbursement for services impacts this significantly.
 |  |
At least two confounders merit consideration in this view of personalised medicine:
1) While it is clear that the identification of a genetic mutation in a given enzyme may significantly impact a patient’s response to a specific drug, the ability to quantify this may be clouded by the biological reality that every enzyme is part of a complex pathway involving multiple enzymes and mutations in these pathway-related enzymes. This may more than compensate for the activity lost (or gained) in enzyme under study;
2) Disease is a process, not a state, meaning that it evolves over time under the influence of environment and lifestyle and patient reaction (interaction) with the disease, itself. Thus, clinical and molecular observations on a patient need to be measured over time, to create a temporal picture of the patient and the disease process. A patient can be represented as a vector, moving through time in a high-dimensional space that is determined by the number of parameters, clinical and molecular, that are being sampled.
In this respect, every patient will be extremely unique and conventional analysis by statistical methods will always be limited in its ability to evaluate and define a “disease” rather than a patient, where n=1. Neither the healthcare system nor the drug development industry are equipped to succeed in a situation where n=1, but an option that provides benefit to both systems and to the patient is one that embraces stratified medicine rather than personalised medicine. In stratified medicine we are looking at how sub-groups of patients may present disease or respond to treatment in a similar manner, within the group, but differing significantly from those in other groups. The basis for this stratification may be of genomic origin but also may reflect true differences in the disease presentation, itself, which have not been adequately identified, evaluated and addressed in clinical practice or in drug research.
Both of these confounders can play a significant role in impacting the potential for “blockbuster drug” development, but reflect the true complexity of the biology that must be addressed, and in manners that reach beyond current systems biology approaches.
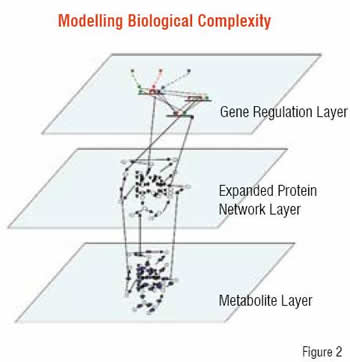
Personalised medicine - Is it more than just genomics?
As noted above, much of personalised medicine has focussed on differences in genetic make-up amongst individuals, as measured by either micro-array technologies, e.g. gene expression, SNPs etc. or most recently, full genome sequencing (e.g. 23andMe, Navigenics, KNOME). While these approaches have provided additional observations on the source of differences amongst individuals, the views that they provide may be too limited, because they are based on application of available, high-throughput technology rather than addressing the true biological complexity that exists. Much of systems biology has evolved from the application of high throughput technologies to provide a more robust “picture” of the patient and of the disease, but while the goal is correct, the approach may be limited to the “looking for keys under the lamp post” syndrome.
The biology of the human system operates in a multi-dimensional, multi-scalar manner ranging from proteins to metabolites, to genes and gene regulation, to differences in time-scales and conditions across multiple tissue types and both macro and microenvironmental differences. While correlations with biological observations may be readily derived with the application of high-throughput technology, a distinct limitation exists in these correlations addressing the mechanistic or causal basis.
Current observations suggest limitations even in the use of Genome Wide Association Studies (GWAS) to yield biomarkers / diagnostics / therapeutic targets specific to a disease and not confounded by the multitude of ancillary differences between individual’s genomes that give rise to the degree of variation exhibited in the human population. In addition, studies also have shown that copy number variation in the genome can not only produce significant associations with disease presentation but also reverse the potential impact of mutations within a single copy of a gene, as evidenced in Craig Venter’s genome.
A more correct representation of biology involves the identification of all its components and the ability to apply the chemical and biological processes as operators on these entities, all integrated to represent the underlying complexity. We are a long way from building an accurate representation of the dynamics of these systems, based on the series of differential equations that can describe the behaviour of each component, but the methodologies exist to enable representation and qualitative reasoning about the systems. This type of modelling, using modelling approaches that have been established in engineering disciplines (e.g. stochastic activity networks, hybrid Petri networks), has the potential to support reasoning with limited data and generate hypotheses leading to rational experimental design and a paradigm that integrates experimental results into the refinement of the models.
Personalised medicine and animal models
As noted above, the limitations in implementing personalised or even stratified medicine into drug development are significant both because of the degree of genomics variation already observed between individuals and the fact that functional biologic variability extends beyond the genome.
This contrasts with the existing variation between species and the dependence on animal models in drug development for toxicity, ADME evaluation and as a model for disease. Significant problems exist in successfully transferring results in animal studies (and even cell lines) to success in humans. Perhaps one of the confounding factors for this problem is the inability to adequately stratify the disease and the patient to enable the effective development of appropriate animal models for the true sub-type of disease or to account for individual variation amongst patients.
Drug safety issues
As noted, a key challenge to the “blockbuster drug” is the impact of drug safety failures. This can be observed in Vioxx, which was a US$ 2.5 billion per year franchise at the time it was pulled from the market and subsequently has led to a US$ 5 billion settlement of legal proceedings from affected patients. An additional impact of the Vioxx situation was Merck’s loss of approximately one-third of its stock value at the time Vioxx was withdrawn (e.g. US$ 44 per share, ~2,100 million shares pre-Vioxx withdrawal, US$ 28 per share, immediately post-withdrawal). The Vioxx lawsuits, numbering 23,800, represented 41,750 plaintiffs, which is likely to be far more than the actual affected population, particularly since the side-effect was only observed in patients on chronic treatment for over 18 months.
Attempts to mitigate drug safety issues have focussed on an ADME / Tox-based perspective, attempting to predict, for a given therapeutic, what the risks are due to its potential for off-target effects. This has emphasised the observation that specificity towards a selected target is difficult to achieve outside of the laboratory because of biological processing, transport, compartmentalisation etc. that comprises the actual physiology in the patient. Obviously differences between individuals at the genomic / proteomic and functional levels (e.g. pathways, networks and processes) all contribute to these problems. A significant opportunity exists that may complete this approach and support identification of potential off-target effects prior to clinical trials.
In a collaborative programme with BIOBASE, Strategic Medicine, Inc. has begun the development and analysis of the complex biological networks occurring at the protein, gene and gene regulation and metabolomics levels, and is building representations of the inter-network connectivity. Using network modelling approaches to support qualitative reasoning about the relative “strengths” of these connections, linkages within the network are evaluated to determine their relative strength, i.e. sensitivity and specificity within the network for producing perturbations to its overall function. The nodes connecting these linkages, which are proteins / enzymes / genes etc. are analysed to identify populations that may be at risk due to polymorphisms or mutations that are prevalent, and then enhanced clinical trial design can proceed to include or exclude such groups from consideration and / or potential labelling of the drug when it proceeds to market, all to lower the overall risk of post-market drug failure.
The goal of this approach is to reduce the failure rate of drugs through the early identification of target-based risk populations and thus enhance their probability for success. This is intended to address some of the current failures presenting in the “blockbuster” model as noted in the earlier part of this article.
Personalised medicine is about biological complexity
This article is intended to present a perspective on whether personalised medicine will drive the end of the blockbuster era for drug development. I have attempted to provide some insights into the current definition of “blockbuster drugs” and how changes to the current model may be achieved in a manner that has both a positive impact on the patient as well as on the economic opportunities for pharmaceutical and biotech development. The key perspective is to recognise both the challenges and the opportunities that the needed changes present to the industry. It may be time to focus on developing a “win-win” strategy that is keen to understand the human physiology and systems thoroughly and thereby enable the successful evolution of “blockbuster drugs” that have long-term, sustainable growth. This can establish the application of personalised medicine for both the patient as well as the pharmaceutical industry in a more systemic and long-reaching manner and produce a highly successful and beneficial synergy.