Rethinking Drug Discovery
Subhadra Dravida, Founder and CEO of Tran-Scell Biologics & TranSTox BioApplications, India.
Prabhat Arya, Department of Organic and Medicinal Chemistry Dr. Reddy's Institute of Life Sciences (DRILS) University of Hyderabad Campus India.
The quest to undertake biological targets that are based on the regulation of signalling pathways is challenging the classical thinking in the drug discovery arena. Historically, the complexity of the biological system was underestimated! It is now well-accepted that compared to old biological targets that were focused on single gene or gene products, we need to undertake targets that are derived from complex and dynamic signaling pathways.
Drug discovery is a tough business! At the same time, it is the domain that provides an opportunity to improve the quality of human health and allows recovering from the suffering due to various biological disorders.
In the past decades, we have witnessed a major limitation in creating the next-generation drugs despite a significant boost in the financial spending for research and development. After the completion of the human genome project, we were guaranteed a plethora of novel biological targets with the hope of producing novel drugs with fewer side effects. The genomics research has made us appreciate the high level complexity of our biological system and at the same time, it challenges the classical thinking in the drug discovery arena.
1.0 Classical drug discovery approach: the pre-genomic era
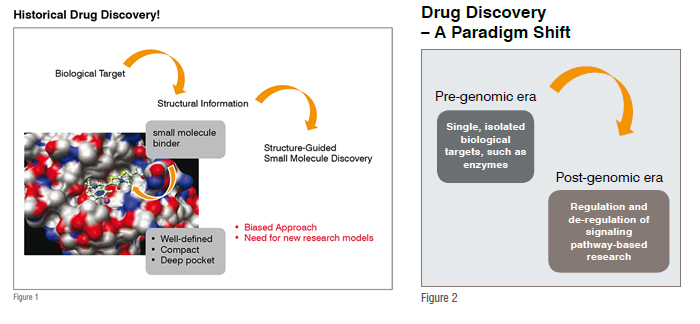
Over the decades, the biomedical drug discovery community’s efforts were focused on a single gene and the gene product, such as isolated enzymes, for example: kinases and phosphatases (See Figure 1). During this course of drug discovery, serious efforts were made to obtain the 3D structural information of a given enzyme; this was then heavily guided by extensive computational studies leading to the design of novel small molecules serving to kick-start the drug discovery programme. In this age, for example, the challenge is to discover novel small molecule that has the potential to hit only one i.e. the desired kinases or phosphatases. The post-genomic era taught us that this is not going to be an easy undertaking, keeping in mind that human genome encodes more than 600 kinases and ~250 phosphatases.
2.0 The post-genomic era
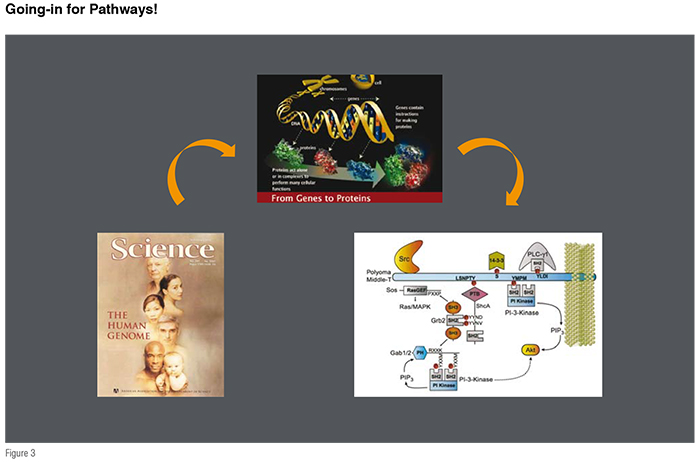
The completion of the human genome that resulted in the indication of nearly 30,000 genes promised a flood of targets to be further undertaken in drug discovery (see Figures 2 and 3). Although highly useful, the information at gene level is not easy to translate to functional protein complexes that are the key to various cell signalling events. Moreover, this information also does not lead to any post-secondary modifications that proteins undergo, such as glycosylation and ubiquitination etc., and, their effects on the signalling functions. In fact significant progress made in genomics and proteomics research has brought us to the doors of a high degree of complexity that lies in our biological system. Furthermore, it also challenges us to develop new research models for understanding the complexity of gene functions, such as the Protein-Protein Interaction (PPI) networks (commonly known as the signalling pathways) that are central to various cell functions in normal as well as disease states. The moment we accept the fact that, proteins do not function in isolation and that they are a part of highly complex network machinery, sets new challenges to examine their role(s) in the drug discovery arena. Why is this case? The pathways that involve multiple protein-protein interactions are highly complex and dynamic. In many cases, even though, we as a community have been successful to obtain the structural information of a given protein-protein interaction, their participation is much more complex (for example, multi-protein complexes), and often leads to a limited information capture for the design of small molecules with desired biological effects.
3.0 Biological targets: signaling pathways-based approach
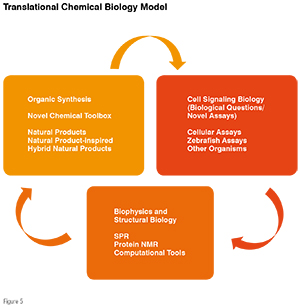
The earlier approaches focused on enzymes (such as kinases and phosphatases) and relied heavily upon the structural information of a given isolated target which would then aid in the design and synthesis of small molecules. With a few exceptions, in most cases, the organic synthesis and medicinal chemistry efforts led to producing "heterocyclic compounds and small molecules that are rich in the sp2 character. The growing desire to undertake biological targets that are focused on protein-protein interactions and on the de-regulation of dynamic signaling pathways is changing our thought process for the choice of small molecules serving as a good starting point to developing the drug discovery research path. Unlike the deep and well-defined pockets that enzymes do offer, in general, PPIs involve a shallow, large surface area with extensive hydrophobic interactions (see Figure 4). Over the years, bioactive natural products have shown an excellent track record as modulators of PPIs, and, in most cases, this is achieved through allosteric sites rather than functioning at the PP-interface. Despite much progress that has been made towards obtaining structural information of a given PPI, due to the nature of these interactions that are generally a part of complex multi-meric protein complex, cell-based screens remain the choice to search for novel small molecules.
The drive to embrace the signalling pathway-based approach is the hallmark in the modern drug discovery arena! And, this need is also seriously questioning our classical approaches to accessing small molecules that were biased towards simple, flat, heterocyclic compounds. A million dollar question in this game, heavily relies on high quality functional cell-based (or commonly called as phenotype or pathway-centric) screening which is the choice of discovering small molecules required in the program. Traditionally, small molecule toolbox within the pharma setting contains compounds that are more biased towards enzymes than on PPIs and pathways. In most cases, these compounds also lack the general features that are commonly found on bioactive natural products, such as 3D architectures, and, rich in chiral display of functional groups etc, functioning on pathways.
4.0 Translational chemical biology: need for new collaborative working models!
There are two prime reasons to explore new research models in drug discovery (see Figure 5), and these are: (i) as discussed above, the traditional pharma collection of small molecules contains compounds that lack the features commonly found in bioactive natural products that are known to function as the modulators of PPIs and signalling pathways; and, (ii) in general, most pharma expertise resides in the classical drug discovery approaches with a focus on kinases, phosphatases and other enzymes. The development of a phenotype or pathway-based screening program is still in its infancy within most or several pharma groups. With an in-depth look at the current limitations covering both points within the pharma community, it is becoming apparent that these bottlenecks can be taken care-of by working closely with the academic groups. Outlined below are some of the advantages that would allow taking care of these two short-comings.
The need for a good starting point as the functional small molecule is the crest of the chemical challenges. The growing interest in undertaking PPI or pathway-based targets is challenging the organic synthesis and medicinal chemistry community to develop novel approaches that allow a rapid generation of small molecules inspired by bioactive natural products or hybrid natural products. The goal here is to build the next generation chemical toolboxes with compounds that have 3D shapes, present sufficient molecular complexity from the medicinal chemistry point of view and are more close to a wide variety of bioactive natural products. To meet these challenges, the academic community has developed several path forward approaches, and, some these include: Diversity-Oriented Synthesis (DOS), Biology-Oriented Synthesis (BIOS) and Functional-Oriented Synthesis (FOS). In all these approaches, the long-term goal is to access a new generation of small molecules that are obtained through the inspiration from bioactive natural products. Because these new synthesis efforts are highly demanding in the methodology development, and, at present, mainly practiced in academic labs, building collaborative working models with the relevant academic community is not surprising. So is the case for the phenotypic or pathway-based cellular screening. From the past several years’ observations, it is now clear that several academic labs across the globe have established an outstanding expertise in this domain that requires working on very high risk projects involving identification and understanding the nature of the biological targets. One of the key features of the translational chemical biology model is that it allows obtaining a thorough understanding of the biological target by an extensive use of biophysics tools, such as, protein NMR, X-Ray, SPR etc.
Exploring some of these demanding and high risk research areas by working closely with the academic community, it is indeed possible to develop next generation path forward approaches within the drug discovery arena. Cities such as Boston, San Diego, Montreal and Toronto have shown tremendous advantages in embracing these new research models. Specifically, Boston leads the way in embracing new research models (for example, Research at the Broad Institute of Harvard and MIT) and, in building a new research culture that has strong roots in academia and the pharma sector. A recent contribution of US$650 million as the philanthropic donation to Broad is the true testimony. It would be nice to see efforts like this in a country like India, who are aspiring to be a powerhouse in building ‘knowledge-based economy’.
5.0 Embracing New Directions: An Example in Cancer Research
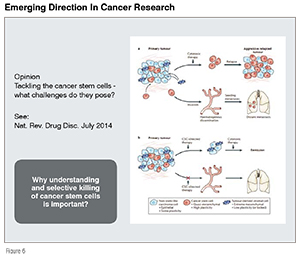
In this section, we outline an example of a translational chemical biology model and its utilisation in embracing a new direction in cancer research (see Figure 6). In addition to the toxicity issues, one of the major problems with cancer drugs is their inability to control the growth/re-birth of cancer cells over a period of time. In the past several years, it has been shown that there is potential to overcome this problem if we are able to design small molecules that are highly selective in killing Cancer Stem Cells (CSCs). The lack of an ability to kill CSCs by most anti-cancer drugs leads to cancer cells formation and this then leads to metastasis. Until very recently, it has been shown that it is possible to identify novel small molecules that are highly effective as selective killers and these compounds in combination with the traditional anti-cancer drugs appear to be a promising path-forward approach. Once again, through developing novel cancer stem cells-based phenotype screens either in cells or in zebrafish combined with a novel chemical toolbox having natural product-inspired and hybrid natural products provides an excellent opportunity to practice working with new models to reach highly unique drug candidates that may prove to be more effective than the traditional anti-cancer drugs.
6.0 Summary
As we have seen for the past several years, the current practice of drug discovery seems to be a losing battle and not much has come out to benefit the society that is desperately looking for next generation effective medicines at an affordable cost. We are hoping that through embracing some of these new research working models and building like-minded teams that are a nice blend of skill-sets from academia and pharma sector would allow reaching the objectives that are not possible to be achieved with classical working models. A challenging task is to build teams to undertake high-risk research programmes, and, this requires a deeper understanding of the need of so called ‘collective competence’. Only time will tell, whether, climbing this mountain would lead to a productive path that the patient community would benefit from, and this remains to be seen in days ahead!
-- Issue 21 --















